This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Caspase-3 Regulatory Mechanisms
From Proteopedia
| Line 1: | Line 1: | ||
| + | <StructureSection load='2h5i_mm1-1.pdb' size='300' side='right' caption='Caspase-3 (PDB entry [[2h5i]])' scene='Sandox_Bay_Serrano/Monomer/1'> | ||
| + | |||
== Introduction == | == Introduction == | ||
| Line 5: | Line 7: | ||
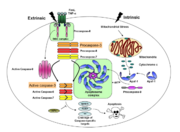
Any apoptotic signal received by the cell causes the activation of initiator caspases (-8 and -9) by associating with other protein platforms to form a functional holoenzyme. These initiator caspases then cleave the executioner caspases -3, -6, and -7. Caspase-3 specifically functions to cleave downstream apoptotic targets as well as both caspase-6 and -7, which in turn cleave their respective targets to induce cell death. Aside from being able to activate caspase-6 and -7, caspase-3 also regulates caspase-9 activity, operating via a feedback loop. This dual action of caspase-3 confers its distinct regulatory mechanisms, resulting in a wider extent of its effects in the apoptotic cascade. | Any apoptotic signal received by the cell causes the activation of initiator caspases (-8 and -9) by associating with other protein platforms to form a functional holoenzyme. These initiator caspases then cleave the executioner caspases -3, -6, and -7. Caspase-3 specifically functions to cleave downstream apoptotic targets as well as both caspase-6 and -7, which in turn cleave their respective targets to induce cell death. Aside from being able to activate caspase-6 and -7, caspase-3 also regulates caspase-9 activity, operating via a feedback loop. This dual action of caspase-3 confers its distinct regulatory mechanisms, resulting in a wider extent of its effects in the apoptotic cascade. | ||
| - | |||
| - | ---- | ||
== Overview of Caspase-3 Structure == | == Overview of Caspase-3 Structure == | ||
| - | |||
| - | <StructureSection load='2h5i_mm1-1.pdb' size='300' side='right' caption='Caspase-3 (PDB entry [[2h5i]])' scene='Sandox_Bay_Serrano/Monomer/1'> | ||
===Dimer Formation === | ===Dimer Formation === | ||
| Line 18: | Line 16: | ||
The active pocket of caspase-3 is defined by <scene name='Sandox_Bay_Serrano/Monomer/2'>Cys-163 and His-121</scene>. Binding of a <scene name='Sandox_Bay_Serrano/Scene01_substrate/4'>substrate</scene>, such as DEVD-CHO to the active site of the enzyme induces a conformational change that allows the L2 and L2' loops to interlock and stabilize the active site <scene name='Sandox_Bay_Serrano/Scene01_substrate/3'></scene>. Like caspase-7, caspase-3 recognizes a Asp-X-X-Asp sequence as a cleavage site in its protein substrates. | The active pocket of caspase-3 is defined by <scene name='Sandox_Bay_Serrano/Monomer/2'>Cys-163 and His-121</scene>. Binding of a <scene name='Sandox_Bay_Serrano/Scene01_substrate/4'>substrate</scene>, such as DEVD-CHO to the active site of the enzyme induces a conformational change that allows the L2 and L2' loops to interlock and stabilize the active site <scene name='Sandox_Bay_Serrano/Scene01_substrate/3'></scene>. Like caspase-7, caspase-3 recognizes a Asp-X-X-Asp sequence as a cleavage site in its protein substrates. | ||
| - | ---- | ||
| - | |||
| - | </StructureSection> | ||
== Caspase-3 Loop Bundle and Active Site== | == Caspase-3 Loop Bundle and Active Site== | ||
| Line 39: | Line 34: | ||
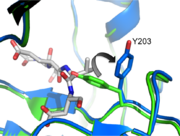
In order to be active and cleave specific apoptotic targets, Caspase-3 must be able to first bind substrate. There are several essential interactions responsible for securing the substrate before cleavage. The binding pocket at <scene name='Caspase-3_Regulatory_Mechanisms/P2/2'>P2</scene> is a hydrophobic patch made up of Y204, W206, and F250 (dark blue residues). This creates a hydrophobic pocket for the P2 residue (in this case, valine), helping it stick to the protein. At <scene name='Caspase-3_Regulatory_Mechanisms/P4/2'>P4</scene> there are contacts that contribute to the specificity of caspase-3. Asparagine 208 hydrogen bonds with an aspartate at P4 along with the backbone nitrogen of F250, creating a preference for a carboxylic acid at the P4 site. | In order to be active and cleave specific apoptotic targets, Caspase-3 must be able to first bind substrate. There are several essential interactions responsible for securing the substrate before cleavage. The binding pocket at <scene name='Caspase-3_Regulatory_Mechanisms/P2/2'>P2</scene> is a hydrophobic patch made up of Y204, W206, and F250 (dark blue residues). This creates a hydrophobic pocket for the P2 residue (in this case, valine), helping it stick to the protein. At <scene name='Caspase-3_Regulatory_Mechanisms/P4/2'>P4</scene> there are contacts that contribute to the specificity of caspase-3. Asparagine 208 hydrogen bonds with an aspartate at P4 along with the backbone nitrogen of F250, creating a preference for a carboxylic acid at the P4 site. | ||
| - | |||
| - | |||
| - | |||
| - | </StructureSection> | ||
== Caspase-3 Regulation== | == Caspase-3 Regulation== | ||
| - | |||
| - | <StructureSection load='1I3O' size='350' side='right' caption='Structure of Caspase-3 with BIR2 domain (PDB entry [[1i3o]])' scene='Caspase-3_Regulatory_Mechanisms/Bir2/1'> | ||
=== Exosite and Allosteric Site=== | === Exosite and Allosteric Site=== | ||
| Line 70: | Line 59: | ||
X-linked inhibitor of apoptosis proteins (XIAP) contains the second baculovirus IAP repeat domain (BIR2) targeting caspase-3 and caspase-7. The <scene name='Caspase-3_Regulatory_Mechanisms/Bir2/2'>BIR2</scene> domain sits directly in the active site of caspase-3 and completely inhibits the protein. The region binding the active site runs in the opposite direction of normal caspase-3 substrates, thus occupying P1 through P4 but avoiding cleavage by the protease. This unique method of inhibition is a critical regulatory mechanism used in cells to control apoptotic caspase activity. | X-linked inhibitor of apoptosis proteins (XIAP) contains the second baculovirus IAP repeat domain (BIR2) targeting caspase-3 and caspase-7. The <scene name='Caspase-3_Regulatory_Mechanisms/Bir2/2'>BIR2</scene> domain sits directly in the active site of caspase-3 and completely inhibits the protein. The region binding the active site runs in the opposite direction of normal caspase-3 substrates, thus occupying P1 through P4 but avoiding cleavage by the protease. This unique method of inhibition is a critical regulatory mechanism used in cells to control apoptotic caspase activity. | ||
| + | </StructureSection> | ||
| + | __NOTOC__ | ||
=References= | =References= | ||
| Line 77: | Line 68: | ||
Hardy, J. A., J. Lam, et al. (2004). "Discovery of an allosteric site in the caspases." Proc Natl Acad Sci U S A 101(34): 12461-12466. | Hardy, J. A., J. Lam, et al. (2004). "Discovery of an allosteric site in the caspases." Proc Natl Acad Sci U S A 101(34): 12461-12466. | ||
| - | |||
| - | </StructureSection> | ||
Revision as of 09:38, 10 July 2013
| |||||||||||
References
Bose, K., C. Pop, et al. (2003). "An uncleavable procaspase-3 mutant has a lower catalytic efficiency but an active site similar to that of mature caspase-3." Biochemistry 42(42): 12298-12310.
Boucher, D., V. Blais, et al. (2012). "Caspase-7 uses an exosite to promote poly(ADP ribose) polymerase 1 proteolysis." Proc Natl Acad Sci U S A 109(15): 5669-5674.
Hardy, J. A., J. Lam, et al. (2004). "Discovery of an allosteric site in the caspases." Proc Natl Acad Sci U S A 101(34): 12461-12466.
Proteopedia Page Contributors and Editors (what is this?)
Scott Eron, Banyuhay P. Serrano, Alexander Berchansky, Yunlong Zhao, Jaime Prilusky, Michal Harel