This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
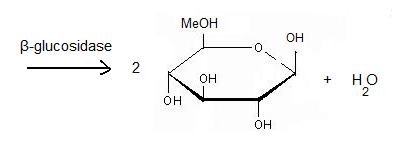
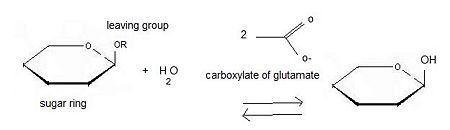
Beta-glucosidase
From Proteopedia
| Line 1: | Line 1: | ||
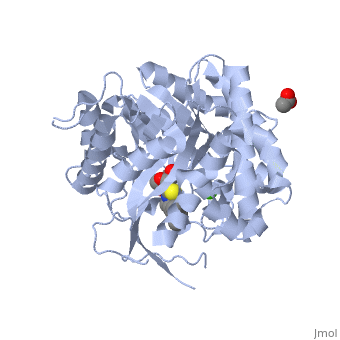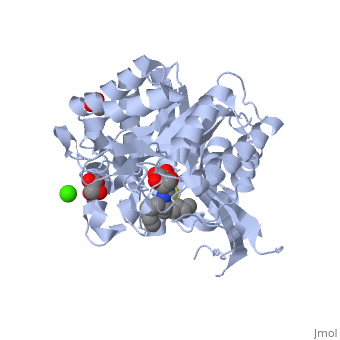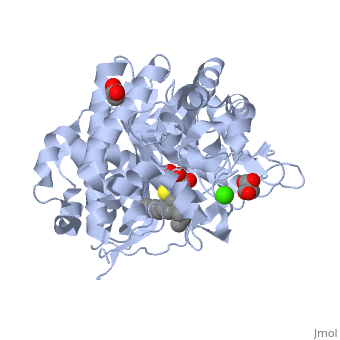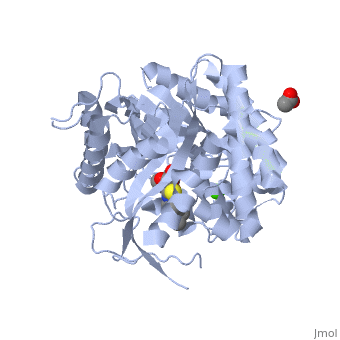
| - | <StructureSection load='2vrj' size='450' side='right' scene='' caption=''> | + | <StructureSection load='2vrj' size='450' side='right' scene='' caption='β-glucosidase complex with calystegine analog, acetate and Ca+2 ion (green) (PDB code [[2vrj]])'> |
[[Image:2vrj.png|left|200px|thumb|Crystal Structure of β-glucosidase, [[2vrj]]]] | [[Image:2vrj.png|left|200px|thumb|Crystal Structure of β-glucosidase, [[2vrj]]]] | ||
{{Clear}} | {{Clear}} | ||
Revision as of 11:37, 3 December 2013
| |||||||||||
3D structures of Beta-glucosidase
Updated on 03-December-2013
3ahx – GBA – Clostridium cellulovorans
3ahy – GB – Trichoderma reesei
3abz – KmGB – Kluyveromyces marxianus
2x40 – TnGB3B – Thermotoga neapolitana
3gno - JrGB residues 38-521 – Japanese rice
2rgl, 2rgm - JrGB residues 29-504
3f4v, 3f5j, 3f5k, 3f5l,3scn, 3sco, 3sct, 3scu, 3scv, 3scw, 3scp, 3scr - JrGB7 (mutant)
3ahz – tGB – termite
3fiy, 3cmj - UbGB catalytic domain (mutant) – Uncultured bacteria
2o9p, 1bga – PpGBB – Paenibacillus polymyxa
2jfe, 3gxd, 3gxi, 3gxm - hGB cystolic – human
3ke0, 3keh - hGB (mutant)
2e3z – PcGB – Phanerochaete crysosporium
2dga - wGB residues 1-520 – wheat
2vff, 1vff – GB – Pyrococcus horikoshii
1oif, 1od0 – TmGB catalytic domain - Thermotoga maritima
1ug6 – GB - Thermus thermophilus
1hxj, 1e1e – ZmGB – Zea mays
1e4l - ZmGB (mutant)
1gon – SsGB – Streptomyces sp.
1qox – GB - Bacillus circulans
1tr1 - BpGBB (mutant) - Bacillus polymyxa
1cbg – GB cyanogenic – White clover
3aiu - rGB residues 50-568 – rye
3f93, 3f94 – PsGB residues 28-840 – Pseudoalteromonas
3f95 – PsGB residues 657-840
3ptk – rGB - rice
3apg – GB – Pyrococcus furiosus
3ta9 – GB – Halothermothrix orenii
3zyz, 3zz1 – GB – Hypocrea jecorina
4hz6 – ubGB – uncultured bacterium
4bce – GB (mutant) – Thermus thermophilus
4iib – AaGB – Aspergillus aculeatus
Beta-glucosidase complex with sugar
3ptm, 3ptq – rGB + glucoside
3ai0 – tGB + glucoside
3ac0 - KmGB + glucoside
2x41, 2x42 - TnGB + glucoside
3air – wGB residues 50-569 + glucoside + dinitrophenol
3ais - wGB residues 50-569 (mutant) + glucoside + aglycone
3aiq - wGB residues 50-569 + aglycone
3aiv - rGB residues 50-568 + aglycone
3aiw - rGB residues 50-568 + glucoside + dinitrophenol
3gnp, 3gnr - JrGB residues 38-521 + glucoside
3aht, 3ahv, 3asq, 3scs – JrGB7 (mutant) + saccharide
1oin - TmGBA + glucoside
3fiz, 3fj0 - UbGB residues 18-482 (mutant) + glucoside
2zox, 2e9l, 2e9m - hGB cystolic + glucoside
2o9s, 2o9r, 2z1s – PpGB + saccharide
2o9t - PpGB + glucoside
1uyq - PpGB (mutant) + glucoside
1bgg - PpGB + gluconate
2jie - BpGB + glucoside
1e4i - BpGB (mutant) + glucoside
2e40 – PcGB + gluconolactone
3vif - NkGB + gluconolactone – Neotermes koshunensis
3vig, 3vii - NkGB + saccharide
3vih – NkGB + glycerol
3vij – NkGB (mutant) + glucose
3vik, 3vil, 3vim, 3vin, 3vio, 3vip – NkGB (mutant) + saccharide
4hz7, 4hz8 - ubGB (mutant) + glucose
1v08 – ZmGB + gluco-tetrazole
1e1f - ZmGB + glucoside
1h49, 1e4n, 1e56 - ZmGB (mutant) + aglycone
1gnx - SsGB + saccharide - Sulfolobus solfataricus
3gfx – hGB + drug
4iic, 4iid, 4iie, 4iif – AaGB + drug
4iig, 4iih – AaGB + saccharide
4i3g – GB + glucose – Streptomyces venezuelae
Beta-glucosidase complex with inhibitor
2wbg, 2wc3, 2wc4, 2vrj, 2jal, 2j75, 2j77, 2j78, 2j79, 2j7b, 2j7d, 2j7e, 2j7f, 2j7g, 2j7h, 2j7c, 2ces, 2cet, 2cbu, 2cbv, 1uz1, 1w3j, 1oim – TmGBA + inhibitor
2cer – SsGB + inhibitor
1e55 - ZmGB (mutant) + inhibitor
3rik, 3ril - hGB + inhibitor
6-phospho-β-glucosidase
1s6y – PGB – Geobacillus stearothermophilus
1up4 – TmPGB
1up6, 1up7 – TmPGB + NAD + G6P
3qom, 4gze – PGB – Lactobacillus plantarum
2xhy – PGB – Escherichia coli
1h4p – PGB I/II – yeast
3eqn – WfPGB – White-rot fungus
3eqo – WfPGB + glucolactone
4b3k, 4b3l – GB – Streptococcus pyogenes
4ipl – SpGB – Streptococcus pneumoniae
4ipn – SpGB + thiocellobiose
Glucan 1,3-β-glucosidase
3n9k – CaGGB + glucoside – Candida albicans
3o6a – CaGGB (mutant)
3ur7, 3ur8 – poGGB – potato
4gzi – poGGB (mutant)
4gzj - poGGB (mutant) + saccharide
Raucaffricine-β-glucosidase
3u57, 3u5u – dpRGB (mutant) – devilpepper
3u5y – dpRGB (mutant)m + secologanin
4a3y, 4atd – seRGB - serpentwood
4atl – seRGB + glucose
4ek7 – seRGB (mutant)
References
- ↑ Aguilar M, Gloster TM, Garcia-Moreno MI, Ortiz Mellet C, Davies GJ, Llebaria A, Casas J, Egido-Gabas M, Garcia Fernandez JM. Molecular basis for beta-glucosidase inhibition by ring-modified calystegine analogues. Chembiochem. 2008 Nov 3;9(16):2612-8. PMID:18833549 doi:10.1002/cbic.200800451
- ↑ http://en.wikipedia.org/wiki/B-glucosidase
- ↑ Davies G, Henrissat B. Structures and mechanisms of glycosyl hydrolases. Structure. 1995 Sep 15;3(9):853-9. PMID:8535779
- ↑ http://www.ebi.ac.uk/interpro/IEntry?ac=IPR018120#PUB00002205
- ↑ http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/CSA/CSA_Site_Wrapper.pl?pdb=2vrj
- ↑ Davies G, Henrissat B. Structures and mechanisms of glycosyl hydrolases. Structure. 1995 Sep 15;3(9):853-9. PMID:8535779
- ↑ http://www.cazy.org/fam/ghf_INV_RET.html#3
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Muriel Breteau, Alexander Berchansky, Joel L. Sussman, David Canner