This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
ATP-dependent DNA ligase from bacteriophage T7
From Proteopedia
(Difference between revisions)
| Line 10: | Line 10: | ||
== Structure == | == Structure == | ||
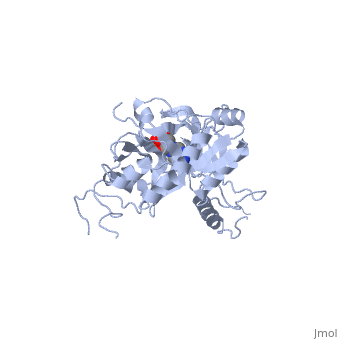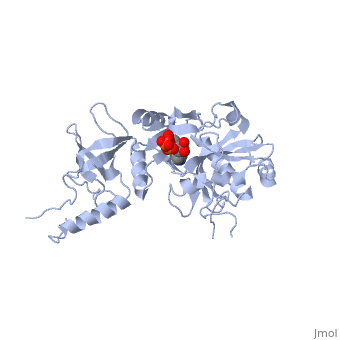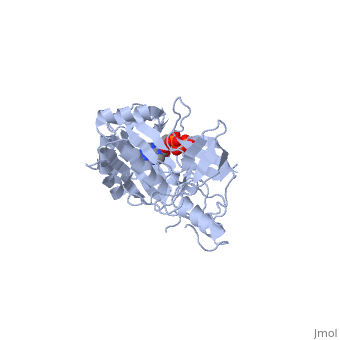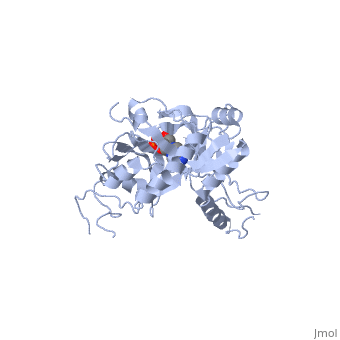
| - | ATP-dependent DNA ligase from bacteriophage T7 is monomeric, forming a tertiary structure consisting of two domains (domain 1 and domain 2). <scene name='56/567310/Domain_1/ | + | ATP-dependent DNA ligase from bacteriophage T7 is monomeric, forming a tertiary structure consisting of two domains (domain 1 and domain 2). <scene name='56/567310/Domain_1/2'>Domain 1</scene> (residues 2:240) contains the ATP binding site. Domain 1 is composed of six alpha helices which surround three antiparallel Beta sheets. |
<scene name='56/567310/Domain_2/1'>Domain 2</scene> (residues 241:349) is composed of an antiparallel Beta sheet and an alpha helix<ref name="Crystal"/>. A groove is formed between the two domains; this groove allows ATP to bind with domain 1. The ribose ring of ATP forms hydrogen bonds with the side chains of <scene name='56/567310/Arg_39_arg_55_glu_93/1'>Arg-39, Arg-55, and Glu-93</scene>. <scene name='56/567310/Lys232_lys238_lys34/2'>Lys-232, Lys-238, and Lys-34</scene> (the catalytic residue) form hydrogen bonds with the three phosphoryl groups of ATP. The 6-amino group of the adenine ring creates hydrogen bonds with the <scene name='56/567310/Ile_33_glu_32/1'>main-chain carbonyl of Ile-33 and the side chain of Glu-32</scene><ref name="Crystal"/>. This could account for the use of ATP rather than GTP. While consisting of 359 residues, residues 121-127, 307-316, and 350-359 are not easily deciphered from the crystalline structure, and are therefore left out of the diagram<ref name="Crystal"/>. Domain 1 contains the N terminus, while domain 2 contains the C terminus. Multiple N and C terminii are shown in the diagram due to the missing residues. | <scene name='56/567310/Domain_2/1'>Domain 2</scene> (residues 241:349) is composed of an antiparallel Beta sheet and an alpha helix<ref name="Crystal"/>. A groove is formed between the two domains; this groove allows ATP to bind with domain 1. The ribose ring of ATP forms hydrogen bonds with the side chains of <scene name='56/567310/Arg_39_arg_55_glu_93/1'>Arg-39, Arg-55, and Glu-93</scene>. <scene name='56/567310/Lys232_lys238_lys34/2'>Lys-232, Lys-238, and Lys-34</scene> (the catalytic residue) form hydrogen bonds with the three phosphoryl groups of ATP. The 6-amino group of the adenine ring creates hydrogen bonds with the <scene name='56/567310/Ile_33_glu_32/1'>main-chain carbonyl of Ile-33 and the side chain of Glu-32</scene><ref name="Crystal"/>. This could account for the use of ATP rather than GTP. While consisting of 359 residues, residues 121-127, 307-316, and 350-359 are not easily deciphered from the crystalline structure, and are therefore left out of the diagram<ref name="Crystal"/>. Domain 1 contains the N terminus, while domain 2 contains the C terminus. Multiple N and C terminii are shown in the diagram due to the missing residues. | ||
Revision as of 04:38, 21 November 2013
ATP-DEPENDENT DNA LIGASE FROM BACTERIOPHAGE T7
| |||||||||||
References
- ↑ McGrath S, van Sinderen D. Bacteriophage: Genetics and Molecular Biology. Norfolk: Caister Academic Press, 2007. Print.
- ↑ Berg, Jeremy M, Stryer, Lubert, Tymoczko, John L. Biochemistry. Sixth edition. New York: W.H. Freeman and Company, 2007: 796. Print.
- ↑ 3.0 3.1 3.2 Doherty AJ, Ashford SR, Subramanya HS, Wigley DB. Bacteriophage T7 DNA ligase. Overexpression, purification, crystallization, and characterization. J Biol Chem. 1996 May 10;271(19):11083-9. PMID:8626651
- ↑ 4.0 4.1 4.2 4.3 4.4 Subramanya HS, Doherty AJ, Ashford SR, Wigley DB. Crystal structure of an ATP-dependent DNA ligase from bacteriophage T7. Cell. 1996 May 17;85(4):607-15. PMID:8653795
Proteopedia Page Contributors and Editors (what is this?)
William Guthrie, Hunter Douglas, Jeremy A. Hammett, Jaime Prilusky