This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 828
From Proteopedia
(Difference between revisions)
| Line 37: | Line 37: | ||
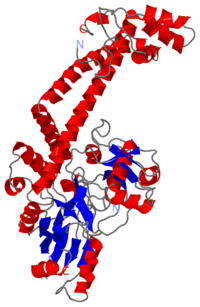
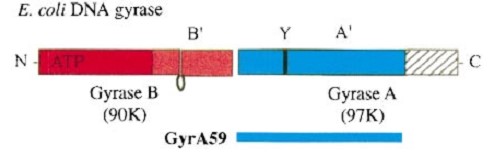
The remaining 30–35 kDa comprising the C-terminal domain (CTD) of GyrA shows a domain forming a b-pinwheel with a positively charged amino-acid perimeter. This carboxy-terminal domain of GyrA (cyan) is required for the introduction of DNA supercoils). | The remaining 30–35 kDa comprising the C-terminal domain (CTD) of GyrA shows a domain forming a b-pinwheel with a positively charged amino-acid perimeter. This carboxy-terminal domain of GyrA (cyan) is required for the introduction of DNA supercoils). | ||
| + | '''The B protein''' has the ATPase domain and the Toprim fold on it. Two ATPase domains dimerize to form a closed conformation. The Toprim fold is a Rossmann fold (beta-alpha-beta-alpha-beta) that contains three invariant acidic residues that coordinate magnesium ions involved in DNA cleavage and DNA religation. | ||
| - | + | ||
| - | The central core of the protein contains a Toprim fold and a DNA-binding core that contains a winged helix domain (WHD), often referred to as a CAP domain. The catalytic tyrosine lies on this WHD. The DNA-binding core consists of the WHD, which leads to a tower domain. . A coiled-coil region leads to a C-terminal domain that forms the main dimer interface | + | '''The central core''' of the protein contains a Toprim fold and a DNA-binding core that contains a winged helix domain (WHD), often referred to as a CAP domain. The catalytic tyrosine lies on this WHD. The DNA-binding core consists of the WHD, which leads to a tower domain. . A coiled-coil region leads to a C-terminal domain that forms the main dimer interface |
the Toprim domain is flexible and that this flexibility can allow the Toprim domain to coordinate with the WHD to form a competent cleavage complex | the Toprim domain is flexible and that this flexibility can allow the Toprim domain to coordinate with the WHD to form a competent cleavage complex | ||
DNA was bent by ~150 degrees through an invariant isoleucine (in topoisomerase II it is I833 and in gyrase it is I172) | DNA was bent by ~150 degrees through an invariant isoleucine (in topoisomerase II it is I833 and in gyrase it is I172) | ||
| Line 45: | Line 46: | ||
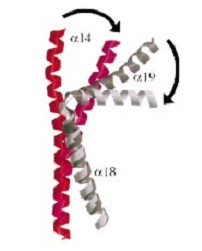
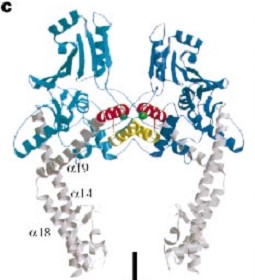
The structures formed a novel beta barrel, which bends DNA by wrapping the nucleic acid around itself. The bending of DNA by gyrase has been proposed as a key mechanism in the ability of gyrase to introduce negative supercoils into the DNA. This is consistent with footprinting data that shows | The structures formed a novel beta barrel, which bends DNA by wrapping the nucleic acid around itself. The bending of DNA by gyrase has been proposed as a key mechanism in the ability of gyrase to introduce negative supercoils into the DNA. This is consistent with footprinting data that shows | ||
that gyrase has a 140-base-pair footprint. Both gyrase and topoisomerase IV CTDs bend DNA, but only gyrase introduces negative supercoils. | that gyrase has a 140-base-pair footprint. Both gyrase and topoisomerase IV CTDs bend DNA, but only gyrase introduces negative supercoils. | ||
| + | |||
| + | |||
| + | ---- | ||
| + | '''Mecanism''' | ||
| + | |||
| + | 40-bp of duplex DNA, the G-segment, bind to the core of the enzyme and are cleaved by the active site tyrosines, while another DNA duplex, the T-segment, is captured through the ATP-induced dimerization of a protein gate, the N-gate. After passage through the transiently broken G-segment (DNA gate), the T-segment exits the protein through another protein gate, the C-gate. ATP hydrolysis and release reset the conformation of the enzyme and DNA to their initial state, poised for another strand-passage event or release of the DNA. | ||
| + | N- and C-gates that open or close to allow T-segment transport through both theprotein and the cleaved G-segment. | ||
| + | |||
---- | ---- | ||
Revision as of 16:20, 24 December 2013
| This Sandbox is Reserved from 06/12/2018, through 30/06/2019 for use in the course "Structural Biology" taught by Bruno Kieffer at the University of Strasbourg, ESBS. This reservation includes Sandbox Reserved 1480 through Sandbox Reserved 1543. |
To get started:
More help: Help:Editing |
| |||||||||||