We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Sandbox reserved 919
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| + | <StructureSection load='3DNM' size='350' frame='true' align='right' caption='Hormone-Sensitive Lipase from [[3dnm]]' scene='58/580297/3dnm_cartoon/2' > | ||
| + | |||
==Introduction to hormone-sensitive lipase== | ==Introduction to hormone-sensitive lipase== | ||
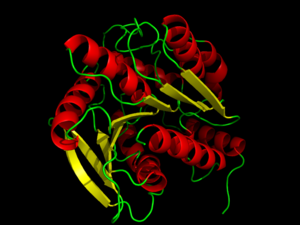
[[Image:HPL_firstimage.png|300 px|left|thumb|Hormone-Sensitive Lipase from [[3dnm]]. Alpha helices and beta sheets are shown in red and yellow, respectively.]] | [[Image:HPL_firstimage.png|300 px|left|thumb|Hormone-Sensitive Lipase from [[3dnm]]. Alpha helices and beta sheets are shown in red and yellow, respectively.]] | ||
| Line 9: | Line 11: | ||
==Structure of hormone-sensitive lipase== | ==Structure of hormone-sensitive lipase== | ||
| - | <StructureSection load='3DNM' size='350' frame='true' align='right' caption='Hormone-Sensitive Lipase from [[3dnm]]' scene='58/580297/3dnm_cartoon/2' > | ||
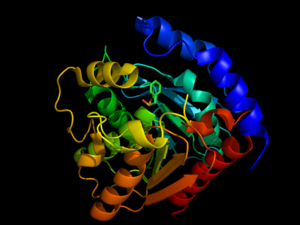
<scene name='58/580297/3dnm_cartoon_dotsribbon/1'>Hormone-sensitive lipases</scene> are generally well-conserved across domains, including prokaryotes, showing 29, 26, and 22% residue overlap in [http://en.wikipedia.org/wiki/Alicyclobacillus ''Alicyclobacillus acidocaldarius''], [http://en.wikipedia.org/wiki/Archaeoglobus ''Archaeoglobus fulgidus''], and [http://en.wikipedia.org/wiki/Bacillus_subtilis ''Bacillus subtilis''], respectively. <ref name="Nam">PMID:19089974</ref> HSL is composed of two main structural domains, consisting of a slightly variable N-terminus (shown in blue in the <scene name='58/580297/3dnm_cartoon/3'>default view</scene>) that is thought to contribute to numerous factors including activity, specificity, regioselectivity, thermophilicity, and thermostability. <ref name="Nam">PMID:19089974</ref> Research speculates that the N-terminal domain, consisting of about 300 residues, mediates protein-protein interactions, and possibly subsequent lipid binding. <ref name= "Yeaman">PMID:14725507</ref> The second domain of HSL is the C-terminal catalytic domain (colors other than blue), which contains serine residue phosphorylation sites as well as the [http://en.wikipedia.org/wiki/Catalytic_triad catalytic triad], viewed <scene name='58/580297/3dnm_triad_zoomedout/1'>here</scene>, a charge relay network that is characteristic of many hydrolases, such as [http://proteopedia.org/wiki/index.php/Chymotrypsin chymotrypsin]. <ref name= "Yeaman">PMID:14725507</ref> With respect to sequence conservation across species, it has been shown that the catalytic domain, including the triad, is conserved across domains, but the domain containing the N-terminus shows little conservation. <ref name="Nam">PMID:19089974</ref> Size-exclusion chromatography studies have shown that HSL has a <scene name='58/580297/3dnm_cartoon_surface/4'>ligand pocket</scene> that is approximately 16Å deep, suggesting that HSL primarily hydrolyzes shorter chained molecules. <ref name="Nam">PMID:19089974</ref> | <scene name='58/580297/3dnm_cartoon_dotsribbon/1'>Hormone-sensitive lipases</scene> are generally well-conserved across domains, including prokaryotes, showing 29, 26, and 22% residue overlap in [http://en.wikipedia.org/wiki/Alicyclobacillus ''Alicyclobacillus acidocaldarius''], [http://en.wikipedia.org/wiki/Archaeoglobus ''Archaeoglobus fulgidus''], and [http://en.wikipedia.org/wiki/Bacillus_subtilis ''Bacillus subtilis''], respectively. <ref name="Nam">PMID:19089974</ref> HSL is composed of two main structural domains, consisting of a slightly variable N-terminus (shown in blue in the <scene name='58/580297/3dnm_cartoon/3'>default view</scene>) that is thought to contribute to numerous factors including activity, specificity, regioselectivity, thermophilicity, and thermostability. <ref name="Nam">PMID:19089974</ref> Research speculates that the N-terminal domain, consisting of about 300 residues, mediates protein-protein interactions, and possibly subsequent lipid binding. <ref name= "Yeaman">PMID:14725507</ref> The second domain of HSL is the C-terminal catalytic domain (colors other than blue), which contains serine residue phosphorylation sites as well as the [http://en.wikipedia.org/wiki/Catalytic_triad catalytic triad], viewed <scene name='58/580297/3dnm_triad_zoomedout/1'>here</scene>, a charge relay network that is characteristic of many hydrolases, such as [http://proteopedia.org/wiki/index.php/Chymotrypsin chymotrypsin]. <ref name= "Yeaman">PMID:14725507</ref> With respect to sequence conservation across species, it has been shown that the catalytic domain, including the triad, is conserved across domains, but the domain containing the N-terminus shows little conservation. <ref name="Nam">PMID:19089974</ref> Size-exclusion chromatography studies have shown that HSL has a <scene name='58/580297/3dnm_cartoon_surface/4'>ligand pocket</scene> that is approximately 16Å deep, suggesting that HSL primarily hydrolyzes shorter chained molecules. <ref name="Nam">PMID:19089974</ref> | ||
Revision as of 17:55, 19 April 2014
| |||||||||||
Additional pages about hormone-sensitive lipase
References
- ↑ 1.0 1.1 Holm C. Molecular mechanisms regulating hormone-sensitive lipase and lipolysis. Biochem Soc Trans. 2003 Dec;31(Pt 6):1120-4. PMID:14641008 doi:http://dx.doi.org/10.1042/
- ↑ 2.0 2.1 Ray H, Beylot M, Arner P, Larrouy D, Langin D, Holm C, Large V. The presence of a catalytically inactive form of hormone-sensitive lipase is associated with decreased lipolysis in abdominal subcutaneous adipose tissue of obese subjects. Diabetes. 2003 Jun;52(6):1417-22. PMID:12765952
- ↑ 3.0 3.1 3.2 3.3 Yeaman SJ. Hormone-sensitive lipase--new roles for an old enzyme. Biochem J. 2004 Apr 1;379(Pt 1):11-22. PMID:14725507 doi:http://dx.doi.org/10.1042/BJ20031811
- ↑ 4.0 4.1 4.2 4.3 Nam KH, Kim MY, Kim SJ, Priyadarshi A, Kwon ST, Koo BS, Yoon SH, Hwang KY. Structural and functional analysis of a novel hormone-sensitive lipase from a metagenome library. Proteins. 2009 Mar;74(4):1036-40. PMID:19089974 doi:http://dx.doi.org/10.1002/prot.22313
- ↑ Kanwar SS, Kaushal RK, Jawed A, Gupta R, Chimni SS. Methods for inhibition of residual lipase activity in colorimetric assay: a comparative study. Indian J Biochem Biophys. 2005 Aug;42(4):233-7. PMID:23923547
- ↑ Kraemer FB, Shen WJ. Hormone-sensitive lipase: control of intracellular tri-(di-)acylglycerol and cholesteryl ester hydrolysis. J Lipid Res. 2002 Oct;43(10):1585-94. PMID:12364542