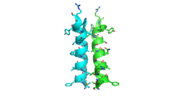This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Brittany Carroll/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 10: | Line 10: | ||
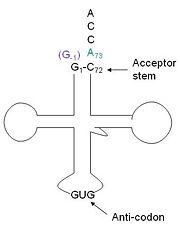
This addition is interesting for multiple reasons. It is one of only a few known reactions where a normal 3’-to-5’ phosphodiester bond is formed in a 3’ –to- 5’ direction. Also, the additional 5’ nucleotide is unique to tRNAHis, with the exception of a tRNAPhe species. Lastly, this modification is essential, at least in yeast. | This addition is interesting for multiple reasons. It is one of only a few known reactions where a normal 3’-to-5’ phosphodiester bond is formed in a 3’ –to- 5’ direction. Also, the additional 5’ nucleotide is unique to tRNAHis, with the exception of a tRNAPhe species. Lastly, this modification is essential, at least in yeast. | ||
| + | |||
== Classification == | == Classification == | ||
| Line 36: | Line 37: | ||
== Structure == | == Structure == | ||
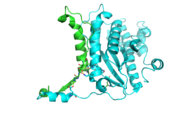
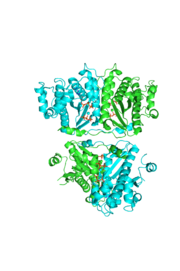
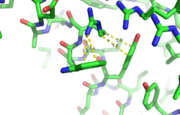
| - | [[Image:thgdimer.png]] | + | [[Image:thgdimer.png|300|left|thumb|The dimer interface is highlighted between monomers, Chain a (green) & Chain b (cyan), showing a large contact area. The two salt bridges between _____ are highlighted as well as some hydrogen bonding. ]] |
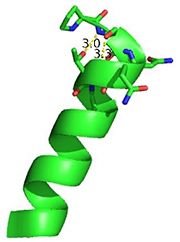
| - | [[Image:alpha_dimer.png]][[Image:tetra.png]] | + | [[Image:alpha_dimer.png|300|right|thumb|The interface between αD of the two monomers.]] |
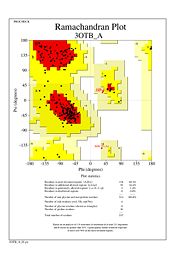
| - | [[Image:ramachandran.jpg]] | + | [[Image:tetra.png|300|left|thumb]] |
| + | [[Image:ramachandran.jpg|300|left|thumb]] | ||
| + | |||
== Structural highlights == | == Structural highlights == | ||
N-terminal helix cap | N-terminal helix cap | ||
Revision as of 14:57, 26 April 2014
tRNA(His) guanylyltransferase
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644