This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Brittany Carroll/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 43: | Line 43: | ||
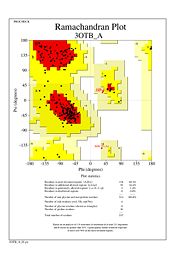
The Ramachandran plot shows that most of the amino acids follow Ramachadran's restraints. The three that are questionable, N32, D67, S75 are all located in turns. | The Ramachandran plot shows that most of the amino acids follow Ramachadran's restraints. The three that are questionable, N32, D67, S75 are all located in turns. | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">color</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | ||
</StructureSection> | </StructureSection> | ||
Revision as of 01:14, 28 April 2014
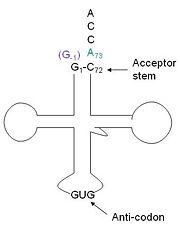
tRNA(His) guanylyltransferase
| |||||||||||
Addition SD Structures of Thg1
3otc, 3otd, 3ote - Thg1 - Homo sapiens
4kgk, 4kgm - Thg1-like - Bacillus thuringiensis
3wbz, 3wc0, 3wc1, 3wc2 - Thg1 - Candida albicans
References
- ↑ Jackman JE, Gott JM, Gray MW. Doing it in reverse: 3'-to-5' polymerization by the Thg1 superfamily. RNA. 2012 May;18(5):886-99. doi: 10.1261/rna.032300.112. Epub 2012 Mar 28. PMID:22456265 doi:http://dx.doi.org/10.1261/rna.032300.112
- ↑ Hyde SJ, Eckenroth BE, Smith BA, Eberley WA, Heintz NH, Jackman JE, Doublie S. tRNAHis guanylyltransferase (THG1), a unique 3'-5' nucleotidyl transferase, shares unexpected structural homology with canonical 5'-3' DNA polymerases. Proc Natl Acad Sci U S A. 2010 Nov 8. PMID:21059936 doi:10.1073/pnas.1010436107
- ↑ Hyde SJ, Rao BS, Eckenroth BE, Jackman JE, Doublie S. Structural Studies of a Bacterial tRNA(HIS) Guanylyltransferase (Thg1)-Like Protein, with Nucleotide in the Activation and Nucleotidyl Transfer Sites. PLoS One. 2013 Jul 3;8(7):e67465. doi: 10.1371/journal.pone.0067465. Print 2013. PMID:23844012 doi:10.1371/journal.pone.0067465
- ↑ Jackman JE, Gott JM, Gray MW. Doing it in reverse: 3'-to-5' polymerization by the Thg1 superfamily. RNA. 2012 May;18(5):886-99. doi: 10.1261/rna.032300.112. Epub 2012 Mar 28. PMID:22456265 doi:http://dx.doi.org/10.1261/rna.032300.112
- ↑ Hyde SJ, Eckenroth BE, Smith BA, Eberley WA, Heintz NH, Jackman JE, Doublie S. tRNAHis guanylyltransferase (THG1), a unique 3'-5' nucleotidyl transferase, shares unexpected structural homology with canonical 5'-3' DNA polymerases. Proc Natl Acad Sci U S A. 2010 Nov 8. PMID:21059936 doi:10.1073/pnas.1010436107
- ↑ Hyde SJ, Eckenroth BE, Smith BA, Eberley WA, Heintz NH, Jackman JE, Doublie S. tRNAHis guanylyltransferase (THG1), a unique 3'-5' nucleotidyl transferase, shares unexpected structural homology with canonical 5'-3' DNA polymerases. Proc Natl Acad Sci U S A. 2010 Nov 8. PMID:21059936 doi:10.1073/pnas.1010436107
- ↑ Hyde SJ, Eckenroth BE, Smith BA, Eberley WA, Heintz NH, Jackman JE, Doublie S. tRNAHis guanylyltransferase (THG1), a unique 3'-5' nucleotidyl transferase, shares unexpected structural homology with canonical 5'-3' DNA polymerases. Proc Natl Acad Sci U S A. 2010 Nov 8. PMID:21059936 doi:10.1073/pnas.1010436107
- ↑ Nakamura A, Nemoto T, Heinemann IU, Yamashita K, Sonoda T, Komoda K, Tanaka I, Soll D, Yao M. Structural basis of reverse nucleotide polymerization. Proc Natl Acad Sci U S A. 2013 Dec 24;110(52):20970-5. doi:, 10.1073/pnas.1321312111. Epub 2013 Dec 9. PMID:24324136 doi:http://dx.doi.org/10.1073/pnas.1321312111
- ↑ http://www.capture.caltech.edu/