This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Jessica Sheehe/Sandbox 1
From Proteopedia
| Line 22: | Line 22: | ||
Dimerization of PKG-Ia is mediated through its leucine zipper motif located at residues 9-44 in the N-terminus of the enzyme (5). The zipper also mediates targeting to substrates leading to their phosphorylation (2). Leucine zippers are right-handed helices that interact through their leucine and isoleucine residues forming a left-handed coiled coil. The primary structure is characterized by a heptad repeat of amino acids (abcdefg) where residues a and d are typically hydrophobic and residues e and g are typically hydrophilic. The structure of the PKG-Ia leucine zipper was solved by NMR in 2005 by Schnell and colleagues (PDB 1ZXA). Using COILS and PARCOILS they predicted the parallel structure of the zipper (6). The helical wheel diagram created using DrawCoil1.0 indicates the residues at the dimer interface. In the corresponding sequence, residues in the a and d positions of the heptad repeat are colored green and the e and g positions are colored purple, indicating the predominant residues at the interface are hydrophobic leucines and isoleucines however the structure is also stabilized by hydrophilic residues in the e and g positions that associate through hydrogen bonding and shield the hydrophobic core. | Dimerization of PKG-Ia is mediated through its leucine zipper motif located at residues 9-44 in the N-terminus of the enzyme (5). The zipper also mediates targeting to substrates leading to their phosphorylation (2). Leucine zippers are right-handed helices that interact through their leucine and isoleucine residues forming a left-handed coiled coil. The primary structure is characterized by a heptad repeat of amino acids (abcdefg) where residues a and d are typically hydrophobic and residues e and g are typically hydrophilic. The structure of the PKG-Ia leucine zipper was solved by NMR in 2005 by Schnell and colleagues (PDB 1ZXA). Using COILS and PARCOILS they predicted the parallel structure of the zipper (6). The helical wheel diagram created using DrawCoil1.0 indicates the residues at the dimer interface. In the corresponding sequence, residues in the a and d positions of the heptad repeat are colored green and the e and g positions are colored purple, indicating the predominant residues at the interface are hydrophobic leucines and isoleucines however the structure is also stabilized by hydrophilic residues in the e and g positions that associate through hydrogen bonding and shield the hydrophobic core. | ||
| + | <table id=branch"Table"> | ||
<td rowspan="2"><[[Image:PKG-Ia Leucine Zipper.png|350px|center|thumb| Leucine Zipper, residues 1-58, PDB 1ZXA]] ><[[Image:PKG-Ia Helical wheel of leucine zipper.png|350px|center|thumb| Helical wheel diagram of PKG-Ia Leucine Zipper, residues 1-58, PDB 1ZXA, made using DrawCoil1.0]]> | <td rowspan="2"><[[Image:PKG-Ia Leucine Zipper.png|350px|center|thumb| Leucine Zipper, residues 1-58, PDB 1ZXA]] ><[[Image:PKG-Ia Helical wheel of leucine zipper.png|350px|center|thumb| Helical wheel diagram of PKG-Ia Leucine Zipper, residues 1-58, PDB 1ZXA, made using DrawCoil1.0]]> | ||
Revision as of 06:07, 28 April 2014
cGMP-dependent protein kinase I alpha (PKG-Ia)
I. Introduction
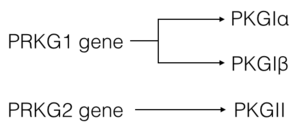
cGMP-dependent protein kinase (PKG) (EC2.7.11.12) is a member of the AGC kinase family that mediates intracellular signaling through serine and threonine phosphorylation. PKG family members are encoded by two genes: PKGI and PKGII. A splice variant in PKGI leads to expression of PKG-Ia and PKG-Iβ (1). Both PKGI isoforms are found in smooth muscle cells, platelets, cerebellum, and hippocampus, however the PKG-Ia isoform is preferentially located in the heart and lungs as well (2). cGMP, produced from guanylate cyclase through activation by natriuretic peptides or NO, is required for full activation of PKG-Ia (1).
II. Isoforms and Homology
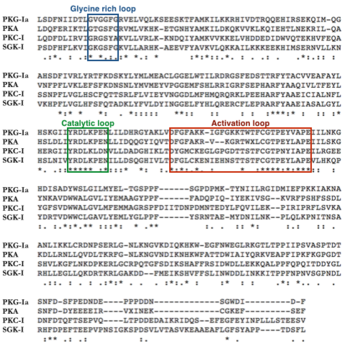
There are two genes that code for PKG—PRKG1 and PRKG2. Alternative splicing of the PRKG1 results in PKG-Ia and PKG-Ib isoforms that differ at their N-terminal region by approximately 65%(1, 3). ClustalW sequence alignment
III. General Structure
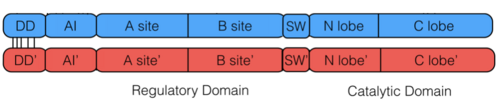
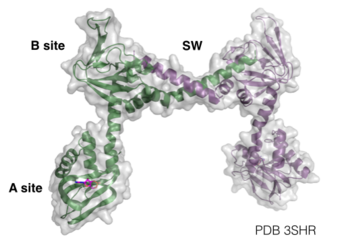
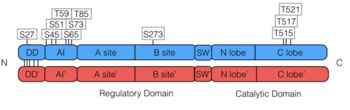
PKG-Ia is a dimer. Each monomer consists of a dimerization domain, an autoinhibitory domain, two cyclic nucleotide binding sites (A site and B site), a switch helical domain, and a catalytic domain (4).
IV. Dimerization Domain
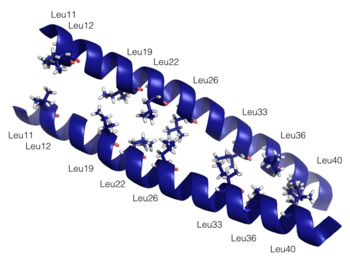
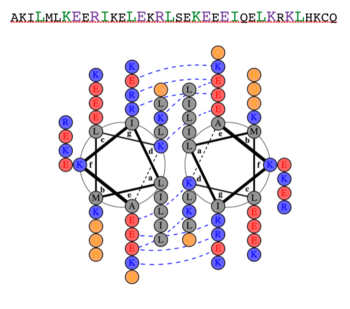
Dimerization of PKG-Ia is mediated through its leucine zipper motif located at residues 9-44 in the N-terminus of the enzyme (5). The zipper also mediates targeting to substrates leading to their phosphorylation (2). Leucine zippers are right-handed helices that interact through their leucine and isoleucine residues forming a left-handed coiled coil. The primary structure is characterized by a heptad repeat of amino acids (abcdefg) where residues a and d are typically hydrophobic and residues e and g are typically hydrophilic. The structure of the PKG-Ia leucine zipper was solved by NMR in 2005 by Schnell and colleagues (PDB 1ZXA). Using COILS and PARCOILS they predicted the parallel structure of the zipper (6). The helical wheel diagram created using DrawCoil1.0 indicates the residues at the dimer interface. In the corresponding sequence, residues in the a and d positions of the heptad repeat are colored green and the e and g positions are colored purple, indicating the predominant residues at the interface are hydrophobic leucines and isoleucines however the structure is also stabilized by hydrophilic residues in the e and g positions that associate through hydrogen bonding and shield the hydrophobic core.