This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Cody Couperus/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 10: | Line 10: | ||
Activity of thrombin is regulated physiologically by the serpin inhibitors: | Activity of thrombin is regulated physiologically by the serpin inhibitors: | ||
| - | * [[Antithrombin]] <ref name="three"/><ref>PMID: 24477356</ref><ref>PMID: 23809129</ref> | + | * [[Antithrombin]] <ref name="three"/><ref name='four'>PMID: 24477356</ref><ref name='five'>PMID: 23809129</ref> |
| - | * Heparin cofactor II <ref name="three"/><ref | + | * Heparin cofactor II <ref name="three"/><ref name='four'/><ref name='five'/> |
| - | * Protein C inhibitor <ref name="three"/><ref | + | * Protein C inhibitor <ref name="three"/><ref name='four'/><ref name='five'/> |
| - | * Protease nexin 1 <ref name="three"/><ref | + | * Protease nexin 1 <ref name="three"/><ref name='four'/><ref name='five'/> |
| - | Generation of thrombin is decreased when thrombin reaches endothelial lining and interacts with [[1fge |thrombomodulin]] which significantly increases activity of thrombin in activating protein C (APC).<ref>PMID: 3029867</ref> This enzyme will go on to inactivate FVa<ref>PMID: 7989361</ref><ref>PMID: 8639840</ref> and [[Factor_VIII | FVIIIa]]<ref | + | Generation of thrombin is decreased when thrombin reaches endothelial lining and interacts with [[1fge |thrombomodulin]] which significantly increases activity of thrombin in activating protein C (APC).<ref>PMID: 3029867</ref> This enzyme will go on to inactivate FVa<ref>PMID: 7989361</ref><ref name='six'>PMID: 8639840</ref> and [[Factor_VIII | FVIIIa]]<ref name='six'/> , cofactors for activation of prothrombin<ref>PMID: 15147718</ref> and [[Factor_Xa | FXa]]<ref>PMID: 10881749</ref>, respectively. |
By balancing substrate specificity, activity, and inhibition thrombin plays a central role in the blood coagulation cascade. <ref name="three"/> | By balancing substrate specificity, activity, and inhibition thrombin plays a central role in the blood coagulation cascade. <ref name="three"/> | ||
| Line 40: | Line 40: | ||
Prothrombin is the zymogen form of thrombin. From N-terminal to C-terminal it consists of a Gla domain, two kringle domains, and a catalytic domain. | Prothrombin is the zymogen form of thrombin. From N-terminal to C-terminal it consists of a Gla domain, two kringle domains, and a catalytic domain. | ||
| - | Prothrombin is activated by prothrombinase which consists of FXa, FVa, calcium, and a phospholipid surface. In vivo the first cleavage occurs at the R320-I321 bond, corresponding to residues 15-16 in thrombin which is the N-terminus of the B chain, producing meizothrombin.<ref>PMID: 22944689</ref> Subsequent cleavage at R271-T272 yields thrombin.<ref | + | Prothrombin is activated by prothrombinase which consists of FXa, FVa, calcium, and a phospholipid surface. In vivo the first cleavage occurs at the R320-I321 bond, corresponding to residues 15-16 in thrombin which is the N-terminus of the B chain, producing meizothrombin.<ref name='seven'>PMID: 22944689</ref> Subsequent cleavage at R271-T272 yields thrombin.<ref name='seven'/> The initial cleavage can also occur at R271 resulting in prothrombin-2 which will then be cleaved at R320 to produce thrombin.<ref>PMID: 1995649</ref> |
| - | After cleavage by prothrombinase the new B chain N-terminus (Ile16) folds into the core protease domain and forms a salt bridge with Asp194.<ref | + | After cleavage by prothrombinase the new B chain N-terminus (Ile16) folds into the core protease domain and forms a salt bridge with Asp194.<ref name='seven'/> This leads to stabilization of regions of the 180s-loop, Na+ binding loop, and γ-loop (zymogen activation domains). These changes provide the correct conformation for the S1 pocket and oxyanion hole for catalysis.<ref name='seven'/><ref>PMID: 15890651</ref> |
| Line 58: | Line 58: | ||
The <scene name='58/583418/B_chain/1' target='0'>B chain</scene> contains the active site of the protein and has numerous notable structural features. The active site is formed at the rims of two interacting 6 stranded <scene name='58/583418/Beta_barrel/2'>beta barrel domains</scene>(N-terminal barrel in red and C-terminal barrel in orange) which are surrounded by 4 helical regions and many turns. | The <scene name='58/583418/B_chain/1' target='0'>B chain</scene> contains the active site of the protein and has numerous notable structural features. The active site is formed at the rims of two interacting 6 stranded <scene name='58/583418/Beta_barrel/2'>beta barrel domains</scene>(N-terminal barrel in red and C-terminal barrel in orange) which are surrounded by 4 helical regions and many turns. | ||
| - | The serine protease <scene name='58/583418/Catalytic_triad/1'>catalytic triad</scene>, based on chymotrypsin numbering, are Ser195, His57, and Asp102. As is common with serine proteases, an <scene name='58/583418/Oxyanion_hole/2'>oxyanion hole</scene> hole is formed by backbone amides of Ser195 and Gly193.<ref | + | The serine protease <scene name='58/583418/Catalytic_triad/1'>catalytic triad</scene>, based on chymotrypsin numbering, are Ser195, His57, and Asp102. As is common with serine proteases, an <scene name='58/583418/Oxyanion_hole/2'>oxyanion hole</scene> hole is formed by backbone amides of Ser195 and Gly193.<ref name='seven'/> This has the functional role of stabilizing the oxyanion intermediate involved in the serine protease mechanism by hydrogen bonding to the oxygen of the P1 residue (traditional substrate-protein nomeclature <ref>PMID: 22925665</ref>. In addition, since thrombin cleaves after Arg/Lys the <scene name='58/583418/S1/2'>S1 specificity site</scene>, formed by the 180s- and 220s- loops, has Asp189 at the base to form a salt bridge with the incoming substrate. Furthermore, the S4 binding pocket accommodates hydrophobic substrate residues. |
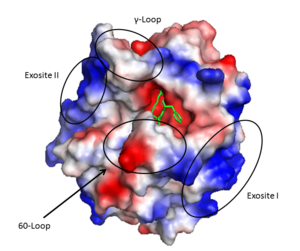
The '''active site''' cleft rims are formed by the hydrophobic and rigid <scene name='58/583418/Loops/2'>60-loops</scene> (residues L60, Y60a, P60b, P60c, W60d, D60e, K60f, N60g, F60h, T60i, and N60g) and the <scene name='58/583418/Loops/1'>γ-loop</scene> (residues T147, W147a, T147b, A147c, N147d, and V147f) while the base is mostly hydrophilic negatively charged amino acids. The cleft is deep compared to more promiscuous serine proteases, consequently substrates must either have a large loop that is cleaved or have favorable interactions with the insertion loops <ref>PMID: 16102053</ref>. | The '''active site''' cleft rims are formed by the hydrophobic and rigid <scene name='58/583418/Loops/2'>60-loops</scene> (residues L60, Y60a, P60b, P60c, W60d, D60e, K60f, N60g, F60h, T60i, and N60g) and the <scene name='58/583418/Loops/1'>γ-loop</scene> (residues T147, W147a, T147b, A147c, N147d, and V147f) while the base is mostly hydrophilic negatively charged amino acids. The cleft is deep compared to more promiscuous serine proteases, consequently substrates must either have a large loop that is cleaved or have favorable interactions with the insertion loops <ref>PMID: 16102053</ref>. | ||
| Line 72: | Line 72: | ||
==Thrombin Allostery== | ==Thrombin Allostery== | ||
| - | Binding of thrombin by sodium or at exosite I stabilizes a form of thrombin that improves substrate recognition.<ref | + | Binding of thrombin by sodium or at exosite I stabilizes a form of thrombin that improves substrate recognition.<ref name='seven'/> This occurs due to energetic linkage between these sites to the S1 binding pocket and oxyanion hole. Rapid kinetic analysis suggests that thrombin is in a dynamic equilibrium that consists of a fast, slow, and inactive state<ref name='seven'/>. There is question as to the physiologic relevance of the inactive state. Regardless, there will be a proportion of fast:slow thrombin and sodium binding to the fast form stabilizes that conformation. Indeed, mutation of the residues involved in sodium binding diminishes the activity of thrombin.<ref name='seven'/> It should be restated, that current data suggest that sodium binding does not induce a conformation change, rather, it stabilizes a conformation of thrombin that has greater activity. |
</StructureSection> | </StructureSection> | ||
Revision as of 05:32, 30 April 2014
| |||||||||||
References
- ↑ Fenton JW 2nd. Thrombin specificity. Ann N Y Acad Sci. 1981;370:468-95. PMID:7023326
- ↑ 2.0 2.1 Coughlin SR. Thrombin signalling and protease-activated receptors. Nature. 2000 Sep 14;407(6801):258-64. PMID:11001069 doi:http://dx.doi.org/10.1038/35025229
- ↑ Crawley JT, Lam JK, Rance JB, Mollica LR, O'Donnell JS, Lane DA. Proteolytic inactivation of ADAMTS13 by thrombin and plasmin. Blood. 2005 Feb 1;105(3):1085-93. Epub 2004 Sep 23. PMID:15388580 doi:http://dx.doi.org/10.1182/blood-2004-03-1101
- ↑ 4.0 4.1 4.2 4.3 4.4 4.5 4.6 Lane DA, Philippou H, Huntington JA. Directing thrombin. Blood. 2005 Oct 15;106(8):2605-12. Epub 2005 Jun 30. PMID:15994286 doi:http://dx.doi.org/10.1182/blood-2005-04-1710
- ↑ Takagi T, Doolittle RF. Amino acid sequence studies on factor XIII and the peptide released during its activation by thrombin. Biochemistry. 1974 Feb 12;13(4):750-6. PMID:4811064
- ↑ Miljic P, Heylen E, Willemse J, Djordjevic V, Radojkovic D, Colovic M, Elezovic I, Hendriks D. Thrombin activatable fibrinolysis inhibitor (TAFI): a molecular link between coagulation and fibrinolysis. Srp Arh Celok Lek. 2010 Jan;138 Suppl 1:74-8. PMID:20229688
- ↑ 7.0 7.1 7.2 7.3 Huntington JA. Natural inhibitors of thrombin. Thromb Haemost. 2014 Apr 1;111(4):583-9. doi: 10.1160/TH13-10-0811. Epub 2014 Jan, 30. PMID:24477356 doi:http://dx.doi.org/10.1160/TH13-10-0811
- ↑ 8.0 8.1 8.2 8.3 Huntington JA. Thrombin inhibition by the serpins. J Thromb Haemost. 2013 Jun;11 Suppl 1:254-64. doi: 10.1111/jth.12252. PMID:23809129 doi:http://dx.doi.org/10.1111/jth.12252
- ↑ Esmon CT. The regulation of natural anticoagulant pathways. Science. 1987 Mar 13;235(4794):1348-52. PMID:3029867
- ↑ Kalafatis M, Rand MD, Mann KG. The mechanism of inactivation of human factor V and human factor Va by activated protein C. J Biol Chem. 1994 Dec 16;269(50):31869-80. PMID:7989361
- ↑ 11.0 11.1 Lu D, Kalafatis M, Mann KG, Long GL. Comparison of activated protein C/protein S-mediated inactivation of human factor VIII and factor V. Blood. 1996 Jun 1;87(11):4708-17. PMID:8639840
- ↑ Duga S, Asselta R, Tenchini ML. Coagulation factor V. Int J Biochem Cell Biol. 2004 Aug;36(8):1393-9. PMID:15147718 doi:http://dx.doi.org/10.1016/j.biocel.2003.08.002
- ↑ Saenko EL, Shima M, Sarafanov AG. Role of activation of the coagulation factor VIII in interaction with vWf, phospholipid, and functioning within the factor Xase complex. Trends Cardiovasc Med. 1999 Oct;9(7):185-92. PMID:10881749
- ↑ 14.0 14.1 14.2 14.3 14.4 14.5 14.6 14.7 Lechtenberg BC, Freund SM, Huntington JA. An ensemble view of thrombin allostery. Biol Chem. 2012 Sep;393(9):889-98. doi: 10.1515/hsz-2012-0178. PMID:22944689 doi:http://dx.doi.org/10.1515/hsz-2012-0178
- ↑ Tijburg PN, van Heerde WL, Leenhouts HM, Hessing M, Bouma BN, de Groot PG. Formation of meizothrombin as intermediate in factor Xa-catalyzed prothrombin activation on endothelial cells. The influence of thrombin on the reaction mechanism. J Biol Chem. 1991 Feb 25;266(6):4017-22. PMID:1995649
- ↑ Bobofchak KM, Pineda AO, Mathews FS, Di Cera E. Energetic and structural consequences of perturbing Gly-193 in the oxyanion hole of serine proteases. J Biol Chem. 2005 Jul 8;280(27):25644-50. Epub 2005 May 12. PMID:15890651 doi:http://dx.doi.org/10.1074/jbc.M503499200
- ↑ Bode W, Mayr I, Baumann U, Huber R, Stone SR, Hofsteenge J. The refined 1.9 A crystal structure of human alpha-thrombin: interaction with D-Phe-Pro-Arg chloromethylketone and significance of the Tyr-Pro-Pro-Trp insertion segment. EMBO J. 1989 Nov;8(11):3467-75. PMID:2583108
- ↑ Page MJ, Di Cera E. Evolution of peptidase diversity. J Biol Chem. 2008 Oct 31;283(44):30010-4. doi: 10.1074/jbc.M804650200. Epub 2008 , Sep 3. PMID:18768474 doi:http://dx.doi.org/10.1074/jbc.M804650200
- ↑ Bode W, Mayr I, Baumann U, Huber R, Stone SR, Hofsteenge J. The refined 1.9 A crystal structure of human alpha-thrombin: interaction with D-Phe-Pro-Arg chloromethylketone and significance of the Tyr-Pro-Pro-Trp insertion segment. EMBO J. 1989 Nov;8(11):3467-75. PMID:2583108
- ↑ Bode W, Mayr I, Baumann U, Huber R, Stone SR, Hofsteenge J. The refined 1.9 A crystal structure of human alpha-thrombin: interaction with D-Phe-Pro-Arg chloromethylketone and significance of the Tyr-Pro-Pro-Trp insertion segment. EMBO J. 1989 Nov;8(11):3467-75. PMID:2583108
- ↑ Bode W, Mayr I, Baumann U, Huber R, Stone SR, Hofsteenge J. The refined 1.9 A crystal structure of human alpha-thrombin: interaction with D-Phe-Pro-Arg chloromethylketone and significance of the Tyr-Pro-Pro-Trp insertion segment. EMBO J. 1989 Nov;8(11):3467-75. PMID:2583108
- ↑ Schechter I, Berger A. On the size of the active site in proteases. I. Papain. 1967. Biochem Biophys Res Commun. 2012 Aug 31;425(3):497-502. doi:, 10.1016/j.bbrc.2012.08.015. PMID:22925665 doi:http://dx.doi.org/10.1016/j.bbrc.2012.08.015
- ↑ Huntington JA. Molecular recognition mechanisms of thrombin. J Thromb Haemost. 2005 Aug;3(8):1861-72. PMID:16102053 doi:http://dx.doi.org/10.1111/j.1538-7836.2005.01363.x
- ↑ Zhang E, Tulinsky A. The molecular environment of the Na+ binding site of thrombin. Biophys Chem. 1997 Jan 31;63(2-3):185-200. PMID:9108691