This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1wco
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
== Structural highlights == | == Structural highlights == | ||
[[1wco]] is a 2 chain structure with sequence from [http://en.wikipedia.org/wiki/Lactococcus_lactis Lactococcus lactis] and [http://en.wikipedia.org/wiki/Monarthropalpus_flavus Monarthropalpus flavus]. This structure supersedes the now removed PDB entry [http://oca.weizmann.ac.il/oca-bin/send-pdb?obs=1&id=1uzt 1uzt]. Full experimental information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1WCO OCA]. <br> | [[1wco]] is a 2 chain structure with sequence from [http://en.wikipedia.org/wiki/Lactococcus_lactis Lactococcus lactis] and [http://en.wikipedia.org/wiki/Monarthropalpus_flavus Monarthropalpus flavus]. This structure supersedes the now removed PDB entry [http://oca.weizmann.ac.il/oca-bin/send-pdb?obs=1&id=1uzt 1uzt]. Full experimental information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1WCO OCA]. <br> | ||
| - | <b>Related:</b> [[1aj1|1aj1]], [[1mqx|1mqx]], [[1mqy|1mqy]], [[1mqz|1mqz]], [[1qow|1qow]], [[1w9n|1w9n]], [[2dde|2dde]], [[2ktn|2ktn]], [[2kto|2kto]]<br> | + | <b>[[Ligand|Ligands:]]</b> <scene name='pdbligand=FPP:FARNESYL+DIPHOSPHATE'>FPP</scene>, <scene name='pdbligand=NAG:N-ACETYL-D-GLUCOSAMINE'>NAG</scene><br> |
| + | <b>[[Non-Standard_Residue|NonStd Res:]]</b> <scene name='pdbligand=DAL:D-ALANINE'>DAL</scene>, <scene name='pdbligand=DBB:D-ALPHA-AMINOBUTYRIC+ACID'>DBB</scene>, <scene name='pdbligand=DBU:(2Z)-2-AMINOBUT-2-ENOIC+ACID'>DBU</scene>, <scene name='pdbligand=DHA:2-AMINO-ACRYLIC+ACID'>DHA</scene>, <scene name='pdbligand=FGA:GAMMA-D-GLUTAMIC+ACID'>FGA</scene>, <scene name='pdbligand=MUB:N-ACETYLMURAMIC+ACID'>MUB</scene><br> | ||
| + | <b>[[Related_structure|Related:]]</b> [[1aj1|1aj1]], [[1mqx|1mqx]], [[1mqy|1mqy]], [[1mqz|1mqz]], [[1qow|1qow]], [[1w9n|1w9n]], [[2dde|2dde]], [[2ktn|2ktn]], [[2kto|2kto]]<br> | ||
<b>Activity:</b> <span class='plainlinks'>[http://en.wikipedia.org/wiki/Glucokinase Glucokinase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=2.7.1.2 2.7.1.2] </span><br> | <b>Activity:</b> <span class='plainlinks'>[http://en.wikipedia.org/wiki/Glucokinase Glucokinase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=2.7.1.2 2.7.1.2] </span><br> | ||
| + | <b>Resources:</b> <span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1wco FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1wco OCA], [http://www.rcsb.org/pdb/explore.do?structureId=1wco RCSB], [http://www.ebi.ac.uk/pdbsum/1wco PDBsum]</span><br> | ||
== Publication Abstract from PubMed == | == Publication Abstract from PubMed == | ||
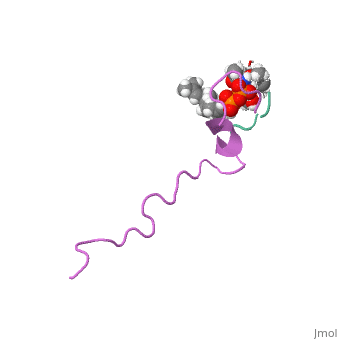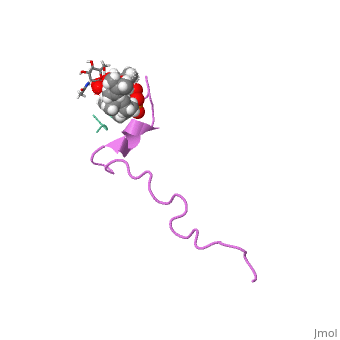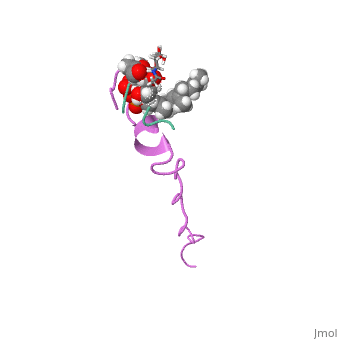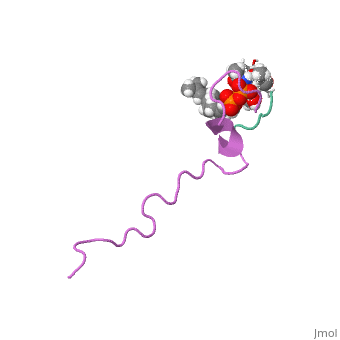
The emerging antibiotics-resistance problem has underlined the urgent need for novel antimicrobial agents. Lantibiotics (lanthionine-containing antibiotics) are promising candidates to alleviate this problem. Nisin, a member of this family, has a unique pore-forming activity against bacteria. It binds to lipid II, the essential precursor of cell wall synthesis. As a result, the membrane permeabilization activity of nisin is increased by three orders of magnitude. Here we report the solution structure of the complex of nisin and lipid II. The structure shows a novel lipid II-binding motif in which the pyrophosphate moiety of lipid II is primarily coordinated by the N-terminal backbone amides of nisin via intermolecular hydrogen bonds. This cage structure provides a rationale for the conservation of the lanthionine rings among several lipid II-binding lantibiotics. The structure of the pyrophosphate cage offers a template for structure-based design of novel antibiotics. | The emerging antibiotics-resistance problem has underlined the urgent need for novel antimicrobial agents. Lantibiotics (lanthionine-containing antibiotics) are promising candidates to alleviate this problem. Nisin, a member of this family, has a unique pore-forming activity against bacteria. It binds to lipid II, the essential precursor of cell wall synthesis. As a result, the membrane permeabilization activity of nisin is increased by three orders of magnitude. Here we report the solution structure of the complex of nisin and lipid II. The structure shows a novel lipid II-binding motif in which the pyrophosphate moiety of lipid II is primarily coordinated by the N-terminal backbone amides of nisin via intermolecular hydrogen bonds. This cage structure provides a rationale for the conservation of the lanthionine rings among several lipid II-binding lantibiotics. The structure of the pyrophosphate cage offers a template for structure-based design of novel antibiotics. | ||
Revision as of 10:02, 30 April 2014
The solution structure of the nisin-lipid II complex
| |||||||||||
Categories: Lactococcus lactis | Monarthropalpus flavus | Bonvin, A M.J J. | Breukink, E. | Hsu, S T.D. | Kaptein, R. | Kruijff, B De. | Lutters, M A.G. | Nuland, N A.J Van. | Tischenko, E. | Antimicrobial | Bacteriocin | Food preservative | Lantibiotic | Peptide-antibiotic complex | Pore formation | Pyrophosphate cage | Thioester