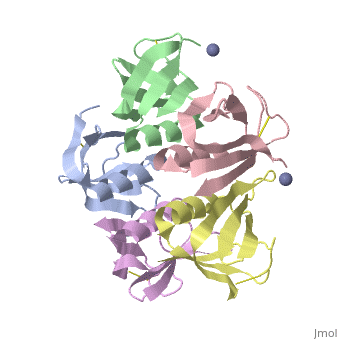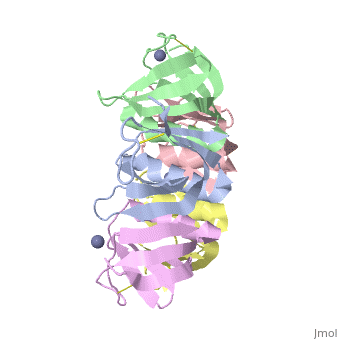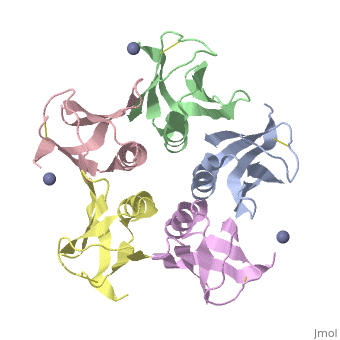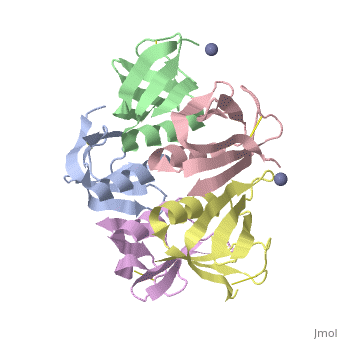This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Shiga toxin
From Proteopedia
| Line 1: | Line 1: | ||
| - | + | <StructureSection load='2xsc' size='450' side='right' scene='' caption=''> | |
==Introduction== | ==Introduction== | ||
| Line 20: | Line 20: | ||
Shiga Toxin acts as an N-glycosidase, removing an adenine from the 28S ribosomal rRNA of a target cell which leads to inhibition of protein elongation and ultimately cellular apoptosis.<ref name=Di>PMID: 21184769</ref> The B subunit is necessary for binding to globo series glycolipid globotriaosylceramide (Gb<sub>3</sub>), a eukaryotic membrane receptor, where it is then endocytosed and proteolytically cleaved into an active A subunit and a B subunit.<ref name=Lenz>PMID: 2170899</ref> The B subunit is not active in the depurination of of 28S rRNA, but is essential for GB<sub>3</sub> binding and therefore essential for toxicity. Once in the cytosol the A subunit is free to interact with and inactivate 28S rRNA. On the A subunit <scene name='Shiga_toxin_1/Active_site_zoomed_in/1'>Tyr77, Tyr114, Glu167, Arg170, and Trp203</scene> are all essential in glycosidic activity.<ref name=Di>PMID: 21184769</ref> This mechanism (B subunit binding to globotriaosylceramide and A subunit depurinating 28S rRNA) is conserved amongst the Stx family as well as the ricin toxin. | Shiga Toxin acts as an N-glycosidase, removing an adenine from the 28S ribosomal rRNA of a target cell which leads to inhibition of protein elongation and ultimately cellular apoptosis.<ref name=Di>PMID: 21184769</ref> The B subunit is necessary for binding to globo series glycolipid globotriaosylceramide (Gb<sub>3</sub>), a eukaryotic membrane receptor, where it is then endocytosed and proteolytically cleaved into an active A subunit and a B subunit.<ref name=Lenz>PMID: 2170899</ref> The B subunit is not active in the depurination of of 28S rRNA, but is essential for GB<sub>3</sub> binding and therefore essential for toxicity. Once in the cytosol the A subunit is free to interact with and inactivate 28S rRNA. On the A subunit <scene name='Shiga_toxin_1/Active_site_zoomed_in/1'>Tyr77, Tyr114, Glu167, Arg170, and Trp203</scene> are all essential in glycosidic activity.<ref name=Di>PMID: 21184769</ref> This mechanism (B subunit binding to globotriaosylceramide and A subunit depurinating 28S rRNA) is conserved amongst the Stx family as well as the ricin toxin. | ||
| - | + | </StructureSection> | |
| + | __NOTOC__ | ||
| + | |||
==3D structures of shiga toxin== | ==3D structures of shiga toxin== | ||
Revision as of 09:29, 3 December 2014
| |||||||||||
3D structures of shiga toxin
Updated on 03-December-2014
1dm0, 1r4q – SdStx α+β chains – Shigella dysenteriae
4m1u – EcStx2 α+β chains + disaccharide – Escherichia coli
Shiga-like toxin 1
1czw, 1czg – pHStx1 β chain (mutant) – phage H30
4ull – pHStx1 β chain – NMR
1c48 - pHStx1 β chain receptor-binding domain (mutant)
1bos, 1qnu – pHStx1 β chain receptor-binding domain + GB3 analog
2xsc – EcStx1 β chain
1d1i, 1cqf – EcStx1 β chain (mutant) + GB3 analog
1c4q, 1d1k – EcStx1 β chain receptor-binding domain (mutant) + GB3 analog
Shiga-like toxin 2
2c5c - pHStx2 β chain + inhibitor
1r4p – EcStx2 α+β chains
3mxg – EcStx2 β chain (mutant)
2bos, 1qoh – EcStx2 β chain receptor-binding domain (mutant) + GB3 analog
2ga4 - Stx2 α+β chains – enterobacteria phage 933W
References
- ↑ 1.0 1.1 Wagner PL, Livny J, Neely MN, Acheson DW, Friedman DI, Waldor MK. Bacteriophage control of Shiga toxin 1 production and release by Escherichia coli. Mol Microbiol. 2002 May;44(4):957-70. PMID:12010491
- ↑ 2.0 2.1 2.2 Herold S, Karch H, Schmidt H. Shiga toxin-encoding bacteriophages--genomes in motion. Int J Med Microbiol. 2004 Sep;294(2-3):115-21. PMID:15493821
- ↑ Asakura H, Makino S, Kobori H, Watarai M, Shirahata T, Ikeda T, Takeshi K. Phylogenetic diversity and similarity of active sites of Shiga toxin (stx) in Shiga toxin-producing Escherichia coli (STEC) isolates from humans and animals. Epidemiol Infect. 2001 Aug;127(1):27-36. PMID:11561972
- ↑ Russell JB, Jarvis GN. Practical mechanisms for interrupting the oral-fecal lifecycle of Escherichia coli. J Mol Microbiol Biotechnol. 2001 Apr;3(2):265-72. PMID:11321582
- ↑ Nishikawa K. Recent progress of Shiga toxin neutralizer for treatment of infections by Shiga toxin-producing Escherichia coli. Arch Immunol Ther Exp (Warsz). 2011 Aug;59(4):239-47. Epub 2011 Jun 5. PMID:21644029 doi:10.1007/s00005-011-0130-5
- ↑ 6.0 6.1 6.2 Fraser ME, Chernaia MM, Kozlov YV, James MN. Crystal structure of the holotoxin from Shigella dysenteriae at 2.5 A resolution. Nat Struct Biol. 1994 Jan;1(1):59-64. PMID:7656009
- ↑ 7.0 7.1 Di R, Kyu E, Shete V, Saidasan H, Kahn PC, Tumer NE. Identification of amino acids critical for the cytotoxicity of Shiga toxin 1 and 2 in Saccharomyces cerevisiae. Toxicon. 2011 Mar 15;57(4):525-39. Epub 2010 Dec 22. PMID:21184769 doi:10.1016/j.toxicon.2010.12.006
- ↑ Roman F, Santa A, Rimanoczky A, Toldi Z, Pataki L. [Isotope study of in vitro K(+) uptake and release of erythrocytes in juvenile diabetes with 86Rb]. Padiatr Grenzgeb. 1990;29(4):339-45. PMID:2170899