This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
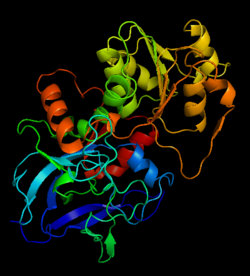
Alcohol dehydrogenase
From Proteopedia
(Difference between revisions)
| Line 109: | Line 109: | ||
**[[2xaa]] – RrADH I + NAD + alcohol<br /> | **[[2xaa]] – RrADH I + NAD + alcohol<br /> | ||
**[[3fx4]] – pADH I + NADP + inhibitor – pig<br /> | **[[3fx4]] – pADH I + NADP + inhibitor – pig<br /> | ||
| + | **[[4w6z]] – yADH I + Zn + NAD derivative<br /> | ||
**[[2w98]], [[2w4q]] – hADH I catalytic domain + NADP + inhibitor<br /> | **[[2w98]], [[2w4q]] – hADH I catalytic domain + NADP + inhibitor<br /> | ||
**[[1hso]] - hADH I α chain + NAD + pyrazole derivative<br /> | **[[1hso]] - hADH I α chain + NAD + pyrazole derivative<br /> | ||
| Line 141: | Line 142: | ||
*ADH IV | *ADH IV | ||
| - | **[[1ye3]], [[8adh]], [[5adh]] - hoADH IV e chain – horse<br /> | + | **[[1ye3]], [[8adh]], [[5adh]], [[4xd2]] - hoADH IV e chain – horse<br /> |
**[[1qlj]] - hoADH IV e chain (mutant) <br /> | **[[1qlj]] - hoADH IV e chain (mutant) <br /> | ||
**[[3iv7]] – ADH IV – ''Corynebacterium glutamicum'' | **[[3iv7]] – ADH IV – ''Corynebacterium glutamicum'' | ||
| Line 177: | Line 178: | ||
**[[1vj0]], [[1vhd]] – TmADH -''Thermotoga maritima''<br /> | **[[1vj0]], [[1vhd]] – TmADH -''Thermotoga maritima''<br /> | ||
**[[2eer]] – ADH – ''Sulfolobus tokodaii''<br /> | **[[2eer]] – ADH – ''Sulfolobus tokodaii''<br /> | ||
| - | **[[3uog]] – ADH – ''Sinorhizobium meliloti'' | + | **[[3uog]] – ADH – ''Sinorhizobium meliloti''<br /> |
| + | **[[4bmn]] – ReADH - ''Ralstonia eutropha''<br /> | ||
*ADH binary complex | *ADH binary complex | ||
| Line 189: | Line 191: | ||
**[[1cdo]] – ADH + NAD - cod<br /> | **[[1cdo]] – ADH + NAD - cod<br /> | ||
**[[1rhc]] – ADH F420-dependent +F420-acetone – ''Methanoculleus thermophilus''<br /> | **[[1rhc]] – ADH F420-dependent +F420-acetone – ''Methanoculleus thermophilus''<br /> | ||
| - | **[[3s2e]] – ReADH + NAD + Zn<br /> | ||
**[[1agn]] – hADH (sigma) +NAD<br /> | **[[1agn]] – hADH (sigma) +NAD<br /> | ||
**[[3pii]] – GsADH + butyramide<br /> | **[[3pii]] – GsADH + butyramide<br /> | ||
**[[3rj5]], [[3rj9]] – SlADH (mutant) + NAD<br /> | **[[3rj5]], [[3rj9]] – SlADH (mutant) + NAD<br /> | ||
| - | **[[3s1l]] – ReADH + Zn | + | **[[3s1l]] – ReADH + Zn <br /> |
**[[3jzd]] – ReADH + NAD<br /> | **[[3jzd]] – ReADH + NAD<br /> | ||
| + | **[[4rqt]] - AtADH P + Zn – ''Arabidopsis thaliana'' <br /> | ||
*ADH ternary complex | *ADH ternary complex | ||
| Line 208: | Line 210: | ||
**[[3s2f]], [[3s2g]] – ReADH + NAD + Zn + furfural<br /> | **[[3s2f]], [[3s2g]] – ReADH + NAD + Zn + furfural<br /> | ||
**[[4gkv]] – ADH + NAD + Zn + peptide – ''Escherichia coli''<br /> | **[[4gkv]] – ADH + NAD + Zn + peptide – ''Escherichia coli''<br /> | ||
| - | **[[4jji]], [[4gl4]], [[3uko]] - AtADH III + NAD + Zn | + | **[[4jji]], [[4gl4]], [[3uko]], [[4rqu]] - AtADH III + NAD + Zn <br /> |
**[[4l0q]] - AtADH III (mutant) + NAD + Zn <br /> | **[[4l0q]] - AtADH III (mutant) + NAD + Zn <br /> | ||
| + | **[[4cpd]] – TtADH + NAD + Zn<br /> | ||
*NADP-dependent ADH | *NADP-dependent ADH | ||
| Line 235: | Line 238: | ||
**[[4jbh]] - PaADH + Zn + Co<br /> | **[[4jbh]] - PaADH + Zn + Co<br /> | ||
**[[4jbi]] - PaADH + NADP + Zn<br /> | **[[4jbi]] - PaADH + NADP + Zn<br /> | ||
| + | **[[4bms]] – ReADH + NADPH<br /> | ||
| + | **[[4bmv]] – ADH + NADPH – ''Sphingobium yanoikuyae''<br /> | ||
*R-specific ADH | *R-specific ADH | ||
Revision as of 11:31, 27 January 2015
| |||||||||||
Additional Resources
For additional information, see: Carbohydrate Metabolism
3D Structures of Alcohol dehydrogenase
Updated on 27-January-2015
References
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol dehydrogenase from Human (Homo sapiens), different isozymes. SCOP. 2009. 1 March 2010 < http://scop.berkeley.edu/data/scop.b.d.c.b.b.c.html>
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Bille V, Remacle J. Simple-kinetic descriptions of alcohol dehydrogenase after immobilization on tresyl-chloride-activated agarose. Eur J Biochem. 1986 Oct 15;160(2):343-8. PMID:3769934
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Blomstrand R, Ostling-Wintzell H, Lof A, McMartin K, Tolf BR, Hedstrom KG. Pyrazoles as inhibitors of alcohol oxidation and as important tools in alcohol research: an approach to therapy against methanol poisoning. Proc Natl Acad Sci U S A. 1979 Jul;76(7):3499-503. PMID:115004
- ↑ Alcohol Dehydrogenase. Worthington Biochemical Corporation . 31 March 2010 < http://http://www.worthington-biochem.com/ADH/default.html>
- ↑ Alcohol Dehydrogenase.Worthington Biochemical Corporation . 31 March 2010 < http://http://www.worthington-biochem.com/ADH/default.html>
- ↑ Goihberg E, Dym O, Tel-Or S, Levin I, Peretz M, Burstein Y. A single proline substitution is critical for the thermostabilization of Clostridium beijerinckii alcohol dehydrogenase. Proteins. 2007 Jan 1;66(1):196-204. PMID:17063493 doi:10.1002/prot.21170
- ↑ Goihberg E, Dym O, Tel-Or S, Shimon L, Frolow F, Peretz M, Burstein Y. Thermal stabilization of the protozoan Entamoeba histolytica alcohol dehydrogenase by a single proline substitution. Proteins. 2008 Feb 7;. PMID:18260103 doi:10.1002/prot.21946
- ↑ Goihberg E, Peretz M, Tel-Or S, Dym O, Shimon L, Frolow F, Burstein Y. Biochemical and Structural Properties of Chimeras Constructed by Exchange of Cofactor-Binding Domains in Alcohol Dehydrogenases from Thermophilic and Mesophilic Microorganisms. Biochemistry. 2010 Feb 9. PMID:20102159 doi:10.1021/bi901730x
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, David Canner, Joel L. Sussman, David Birrer