This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Aminopeptidase
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
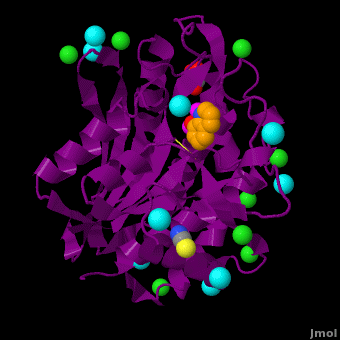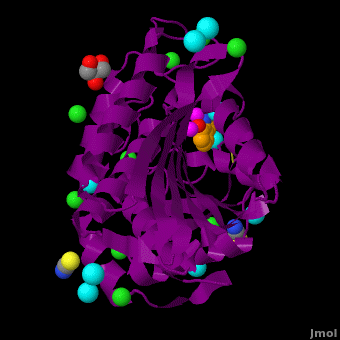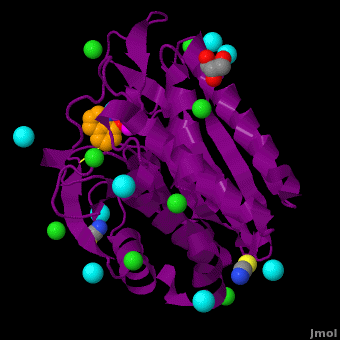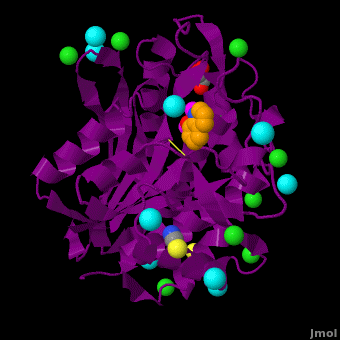
| - | <StructureSection load='VH9.pdb' size='450' side='right' scene='Journal:JBIC:15/Cv/1' caption='Bacterial leucine aminopeptidase complex with 8-hydroxyquinoline, glycerol, SCN, Zn+2 (magenta), Na+ (cyan) and Cl- (green) ions (PDB code [[3vh9]])'>[[Aminopeptidase|Aminopeptidases]] (AP) ([[EC]] 3) are metal - mostly Zn-dependent enzymes involved in the digestion of proteins. Cytosol AP (Cyt-AP), deblocking AP (DAP) and AP N (APN) remove N-terminal amino acids. The AP are classified by the amino acid which they hydrolyze. Other types of AP are: Cold-activated AP (Col-AP) | + | <StructureSection load='VH9.pdb' size='450' side='right' scene='Journal:JBIC:15/Cv/1' caption='Bacterial leucine aminopeptidase complex with 8-hydroxyquinoline, glycerol, SCN, Zn+2 (magenta), Na+ (cyan) and Cl- (green) ions (PDB code [[3vh9]])'>[[Aminopeptidase|Aminopeptidases]] (AP) ([[EC]] 3) are metal - mostly Zn-dependent enzymes involved in the digestion of proteins. Cytosol AP (Cyt-AP), deblocking AP (DAP) and AP N (APN) remove N-terminal amino acids. The AP are classified by the amino acid which they hydrolyze. Other types of AP are:<br /> |
| + | * '''Cold-activated AP''' (Col-AP)<br /> | ||
| + | * '''Heat stable AP''' from ''Thermus thermophilus'' (AmpT)<br /> | ||
| + | * '''AP from ''Staphylococcus aureus''''' (AmpS)<br /> | ||
| + | * '''SGAP''' from ''Stereomyces griseus''. See details in [[Streptomyces griseus Aminopeptidase (SGAP)]].<br /> | ||
| + | * '''Alanine aminopeptidase''' is called '''aminopeptidase N'''. See details in [[Aminopeptidase N]].<br '> | ||
| + | Aminopeptidases catalyze a release of an N-terminal amino acid from a peptide, amide, or arylamide. Beta-peptidyl AP (BapA) cleaves N-terminal β-homoamino acid from peptides of length 2 to 6. | ||
== Aminopeptidase from ''Aeromonas proteolytica''<ref>DOI 10.1007/s00775-012-0873-4</ref> == | == Aminopeptidase from ''Aeromonas proteolytica''<ref>DOI 10.1007/s00775-012-0873-4</ref> == | ||
Revision as of 09:20, 22 December 2014
| |||||||||||
3D Structures of Aminopeptidase
Updated on 22-December-2014
Additional Resources
For additional information, see:
Amino Acid Synthesis & Metabolism
Streptomyces griseus Aminopeptidase (SGAP)
References
- ↑ Hanaya K, Suetsugu M, Saijo S, Yamato I, Aoki S. Potent inhibition of dinuclear zinc(II) peptidase, an aminopeptidase from Aeromonas proteolytica, by 8-quinolinol derivatives: inhibitor design based on Zn(2+) fluorophores, kinetic, and X-ray crystallographic study. J Biol Inorg Chem. 2012 Feb 5. PMID:22311113 doi:10.1007/s00775-012-0873-4
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, David Canner, Joel L. Sussman, Eran Hodis