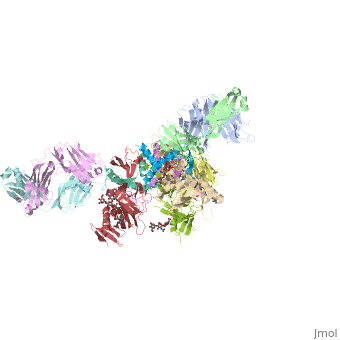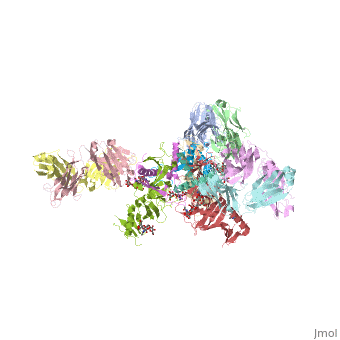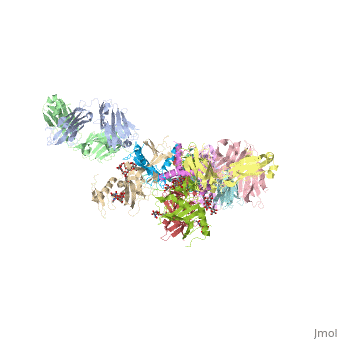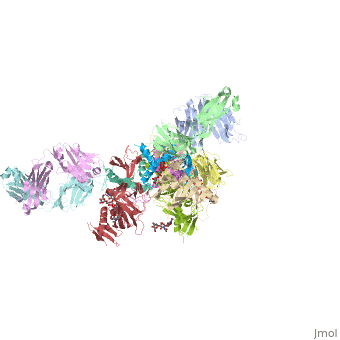
| Structural highlights
Function
[VGP_EBOZ5] GP1 is responsible for binding to the receptor(s) on target cells. Interacts with CD209/DC-SIGN and CLEC4M/DC-SIGNR which act as cofactors for virus entry into the host cell. Binding to CD209 and CLEC4M, which are respectively found on dendritic cells (DCs), and on endothelial cells of liver sinusoids and lymph node sinuses, facilitate infection of macrophages and endothelial cells. These interactions not only facilitate virus cell entry, but also allow capture of viral particles by DCs and subsequent transmission to susceptible cells without DCs infection (trans infection). Binding to the macrophage specific lectin CLEC10A also seems to enhance virus infectivity. Interaction with FOLR1/folate receptor alpha may be a cofactor for virus entry in some cell types, although results are contradictory. Members of the Tyro3 receptor tyrosine kinase family also seem to be cell entry factors in filovirus infection. Once attached, the virions are internalized through clathrin-dependent endocytosis and/or macropinocytosis. After internalization of the virus into the endosomes of the host cell, proteolysis of GP1 by two cysteine proteases, CTSB/cathepsin B and CTSL/cathepsin L presumably induces a conformational change of GP2, unmasking its fusion peptide and initiating membranes fusion (By similarity). GP2 acts as a class I viral fusion protein. Under the current model, the protein has at least 3 conformational states: pre-fusion native state, pre-hairpin intermediate state, and post-fusion hairpin state. During viral and target cell membrane fusion, the coiled coil regions (heptad repeats) assume a trimer-of-hairpins structure, positioning the fusion peptide in close proximity to the C-terminal region of the ectodomain. The formation of this structure appears to drive apposition and subsequent fusion of viral and target cell membranes. Responsible for penetration of the virus into the cell cytoplasm by mediating the fusion of the membrane of the endocytosed virus particle with the endosomal membrane. Low pH in endosomes induces an irreversible conformational change in GP2, releasing the fusion hydrophobic peptide (By similarity). GP1,2 mediates endothelial cell activation and decreases endothelial barrier function. Mediates activation of primary macrophages. At terminal stages of the viral infection, when its expression is high, GP1,2 down-modulates the expression of various host cell surface molecules that are essential for immune surveillance and cell adhesion. Down-modulates integrins ITGA1, ITGA2, ITGA3, ITGA4, ITGA5, ITGA6, ITGAV and ITGB1. GP1,2 alters the cellular recycling of the dimer alpha-V/beta-3 via a dynamin-dependent pathway. Decrease in the host cell surface expression of various adhesion molecules may lead to cell detachment, contributing to the disruption of blood vessel integrity and hemorrhages developed during Ebola virus infection (cytotoxicity). This cytotoxicity appears late in the infection, only after the massive release of viral particles by infected cells. Down-modulation of host MHC-I, leading to altered recognition by immune cells, may explain the immune suppression and inflammatory dysfunction linked to Ebola infection. Also down-modulates EGFR surface expression (By similarity). GP2delta is part of the complex GP1,2delta released by host ADAM17 metalloprotease. This secreted complex may play a role in the pathogenesis of the virus by efficiently blocking the neutralizing antibodies that would otherwise neutralize the virus surface glycoproteins GP1,2. Might therefore contribute to the lack of inflammatory reaction seen during infection in spite the of extensive necrosis and massive virus production. GP1,2delta does not seem to be involved in activation of primary macrophages (By similarity). [VGP_EBOZM] GP1 is responsible for binding to the receptor(s) on target cells. Interacts with CD209/DC-SIGN and CLEC4M/DC-SIGNR which act as cofactors for virus entry into the host cell. Binding to CD209 and CLEC4M, which are respectively found on dendritic cells (DCs), and on endothelial cells of liver sinusoids and lymph node sinuses, facilitate infection of macrophages and endothelial cells. These interactions not only facilitate virus cell entry, but also allow capture of viral particles by DCs and subsequent transmission to susceptible cells without DCs infection (trans infection). Binding to the macrophage specific lectin CLEC10A also seem to enhance virus infectivity. Interaction with FOLR1/folate receptor alpha may be a cofactor for virus entry in some cell types, although results are contradictory. Members of the Tyro3 receptor tyrosine kinase family also seem to be cell entry factors in filovirus infection. Once attached, the virions are internalized through clathrin-dependent endocytosis and/or macropinocytosis. After internalization of the virus into the endosomes of the host cell, proteolysis of GP1 by two cysteine proteases, CTSB/cathepsin B and CTSL/cathepsin L presumably induces a conformational change of GP2, unmasking its fusion peptide and initiating membranes fusion.[1] [2] [3] [4] [5] [6] [7] [8] GP2 acts as a class I viral fusion protein. Under the current model, the protein has at least 3 conformational states: pre-fusion native state, pre-hairpin intermediate state, and post-fusion hairpin state. During viral and target cell membrane fusion, the coiled coil regions (heptad repeats) assume a trimer-of-hairpins structure, positioning the fusion peptide in close proximity to the C-terminal region of the ectodomain. The formation of this structure appears to drive apposition and subsequent fusion of viral and target cell membranes. Responsible for penetration of the virus into the cell cytoplasm by mediating the fusion of the membrane of the endocytosed virus particle with the endosomal membrane. Low pH in endosomes induces an irreversible conformational change in GP2, releasing the fusion hydrophobic peptide.[9] [10] [11] [12] [13] [14] [15] [16] GP1,2 mediates endothelial cell activation and decreases endothelial barrier function. Mediates activation of primary macrophages. At terminal stages of the viral infection, when its expression is high, GP1,2 down-modulates the expression of various host cell surface molecules that are essential for immune surveillance and cell adhesion. Down-modulates integrins ITGA1, ITGA2, ITGA3, ITGA4, ITGA5, ITGA6, ITGAV and ITGB1. GP1,2 alters the cellular recycling of the dimer alpha-V/beta-3 via a dynamin-dependent pathway. Decrease in the host cell surface expression of various adhesion molecules may lead to cell detachment, contributing to the disruption of blood vessel integrity and hemorrhages developed during Ebola virus infection (cytotoxicity). This cytotoxicity appears late in the infection, only after the massive release of viral particles by infected cells. Down-modulation of host MHC-I, leading to altered recognition by immune cells, may explain the immune suppression and inflammatory dysfunction linked to Ebola infection. Also down-modulates EGFR surface expression.[17] [18] [19] [20] [21] [22] [23] [24] GP2delta is part of the complex GP1,2delta released by host ADAM17 metalloprotease. This secreted complex may play a role in the pathogenesis of the virus by efficiently blocking the neutralizing antibodies that would otherwise neutralize the virus surface glycoproteins GP1,2. Might therefore contribute to the lack of inflammatory reaction seen during infection in spite the of extensive necrosis and massive virus production. GP1,2delta does not seem to be involved in activation of primary macrophages.[25] [26] [27] [28] [29] [30] [31] [32]
Evolutionary Conservation
Check, as determined by ConSurfDB. You may read the explanation of the method and the full data available from ConSurf.
Publication Abstract from PubMed
Ebola virus (EBOV) entry requires the surface glycoprotein (GP) to initiate attachment and fusion of viral and host membranes. Here we report the crystal structure of EBOV GP in its trimeric, pre-fusion conformation (GP1+GP2) bound to a neutralizing antibody, KZ52, derived from a human survivor of the 1995 Kikwit outbreak. Three GP1 viral attachment subunits assemble to form a chalice, cradled by the GP2 fusion subunits, while a novel glycan cap and projected mucin-like domain restrict access to the conserved receptor-binding site sequestered in the chalice bowl. The glycocalyx surrounding GP is likely central to immune evasion and may explain why survivors have insignificant neutralizing antibody titres. KZ52 recognizes a protein epitope at the chalice base where it clamps several regions of the pre-fusion GP2 to the amino terminus of GP1. This structure provides a template for unravelling the mechanism of EBOV GP-mediated fusion and for future immunotherapeutic development.
Structure of the Ebola virus glycoprotein bound to an antibody from a human survivor.,Lee JE, Fusco ML, Hessell AJ, Oswald WB, Burton DR, Saphire EO Nature. 2008 Jul 10;454(7201):177-82. PMID:18615077[33]
From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.
See Also
References
- ↑ Yang ZY, Duckers HJ, Sullivan NJ, Sanchez A, Nabel EG, Nabel GJ. Identification of the Ebola virus glycoprotein as the main viral determinant of vascular cell cytotoxicity and injury. Nat Med. 2000 Aug;6(8):886-9. PMID:10932225 doi:10.1038/78645
- ↑ Alvarez CP, Lasala F, Carrillo J, Muniz O, Corbi AL, Delgado R. C-type lectins DC-SIGN and L-SIGN mediate cellular entry by Ebola virus in cis and in trans. J Virol. 2002 Jul;76(13):6841-4. PMID:12050398
- ↑ Wahl-Jensen VM, Afanasieva TA, Seebach J, Stroher U, Feldmann H, Schnittler HJ. Effects of Ebola virus glycoproteins on endothelial cell activation and barrier function. J Virol. 2005 Aug;79(16):10442-50. PMID:16051836 doi:79/16/10442
- ↑ Wahl-Jensen V, Kurz SK, Hazelton PR, Schnittler HJ, Stroher U, Burton DR, Feldmann H. Role of Ebola virus secreted glycoproteins and virus-like particles in activation of human macrophages. J Virol. 2005 Feb;79(4):2413-9. PMID:15681442 doi:79/4/2413
- ↑ Alazard-Dany N, Volchkova V, Reynard O, Carbonnelle C, Dolnik O, Ottmann M, Khromykh A, Volchkov VE. Ebola virus glycoprotein GP is not cytotoxic when expressed constitutively at a moderate level. J Gen Virol. 2006 May;87(Pt 5):1247-57. PMID:16603527 doi:87/5/1247
- ↑ Marzi A, Akhavan A, Simmons G, Gramberg T, Hofmann H, Bates P, Lingappa VR, Pohlmann S. The signal peptide of the ebolavirus glycoprotein influences interaction with the cellular lectins DC-SIGN and DC-SIGNR. J Virol. 2006 Jul;80(13):6305-17. PMID:16775318 doi:10.1128/JVI.02545-05
- ↑ Saeed MF, Kolokoltsov AA, Albrecht T, Davey RA. Cellular entry of ebola virus involves uptake by a macropinocytosis-like mechanism and subsequent trafficking through early and late endosomes. PLoS Pathog. 2010 Sep 16;6(9):e1001110. doi: 10.1371/journal.ppat.1001110. PMID:20862315 doi:10.1371/journal.ppat.1001110
- ↑ Bhattacharyya S, Warfield KL, Ruthel G, Bavari S, Aman MJ, Hope TJ. Ebola virus uses clathrin-mediated endocytosis as an entry pathway. Virology. 2010 May 25;401(1):18-28. doi: 10.1016/j.virol.2010.02.015. Epub 2010, Mar 3. PMID:20202662 doi:10.1016/j.virol.2010.02.015
- ↑ Yang ZY, Duckers HJ, Sullivan NJ, Sanchez A, Nabel EG, Nabel GJ. Identification of the Ebola virus glycoprotein as the main viral determinant of vascular cell cytotoxicity and injury. Nat Med. 2000 Aug;6(8):886-9. PMID:10932225 doi:10.1038/78645
- ↑ Alvarez CP, Lasala F, Carrillo J, Muniz O, Corbi AL, Delgado R. C-type lectins DC-SIGN and L-SIGN mediate cellular entry by Ebola virus in cis and in trans. J Virol. 2002 Jul;76(13):6841-4. PMID:12050398
- ↑ Wahl-Jensen VM, Afanasieva TA, Seebach J, Stroher U, Feldmann H, Schnittler HJ. Effects of Ebola virus glycoproteins on endothelial cell activation and barrier function. J Virol. 2005 Aug;79(16):10442-50. PMID:16051836 doi:79/16/10442
- ↑ Wahl-Jensen V, Kurz SK, Hazelton PR, Schnittler HJ, Stroher U, Burton DR, Feldmann H. Role of Ebola virus secreted glycoproteins and virus-like particles in activation of human macrophages. J Virol. 2005 Feb;79(4):2413-9. PMID:15681442 doi:79/4/2413
- ↑ Alazard-Dany N, Volchkova V, Reynard O, Carbonnelle C, Dolnik O, Ottmann M, Khromykh A, Volchkov VE. Ebola virus glycoprotein GP is not cytotoxic when expressed constitutively at a moderate level. J Gen Virol. 2006 May;87(Pt 5):1247-57. PMID:16603527 doi:87/5/1247
- ↑ Marzi A, Akhavan A, Simmons G, Gramberg T, Hofmann H, Bates P, Lingappa VR, Pohlmann S. The signal peptide of the ebolavirus glycoprotein influences interaction with the cellular lectins DC-SIGN and DC-SIGNR. J Virol. 2006 Jul;80(13):6305-17. PMID:16775318 doi:10.1128/JVI.02545-05
- ↑ Saeed MF, Kolokoltsov AA, Albrecht T, Davey RA. Cellular entry of ebola virus involves uptake by a macropinocytosis-like mechanism and subsequent trafficking through early and late endosomes. PLoS Pathog. 2010 Sep 16;6(9):e1001110. doi: 10.1371/journal.ppat.1001110. PMID:20862315 doi:10.1371/journal.ppat.1001110
- ↑ Bhattacharyya S, Warfield KL, Ruthel G, Bavari S, Aman MJ, Hope TJ. Ebola virus uses clathrin-mediated endocytosis as an entry pathway. Virology. 2010 May 25;401(1):18-28. doi: 10.1016/j.virol.2010.02.015. Epub 2010, Mar 3. PMID:20202662 doi:10.1016/j.virol.2010.02.015
- ↑ Yang ZY, Duckers HJ, Sullivan NJ, Sanchez A, Nabel EG, Nabel GJ. Identification of the Ebola virus glycoprotein as the main viral determinant of vascular cell cytotoxicity and injury. Nat Med. 2000 Aug;6(8):886-9. PMID:10932225 doi:10.1038/78645
- ↑ Alvarez CP, Lasala F, Carrillo J, Muniz O, Corbi AL, Delgado R. C-type lectins DC-SIGN and L-SIGN mediate cellular entry by Ebola virus in cis and in trans. J Virol. 2002 Jul;76(13):6841-4. PMID:12050398
- ↑ Wahl-Jensen VM, Afanasieva TA, Seebach J, Stroher U, Feldmann H, Schnittler HJ. Effects of Ebola virus glycoproteins on endothelial cell activation and barrier function. J Virol. 2005 Aug;79(16):10442-50. PMID:16051836 doi:79/16/10442
- ↑ Wahl-Jensen V, Kurz SK, Hazelton PR, Schnittler HJ, Stroher U, Burton DR, Feldmann H. Role of Ebola virus secreted glycoproteins and virus-like particles in activation of human macrophages. J Virol. 2005 Feb;79(4):2413-9. PMID:15681442 doi:79/4/2413
- ↑ Alazard-Dany N, Volchkova V, Reynard O, Carbonnelle C, Dolnik O, Ottmann M, Khromykh A, Volchkov VE. Ebola virus glycoprotein GP is not cytotoxic when expressed constitutively at a moderate level. J Gen Virol. 2006 May;87(Pt 5):1247-57. PMID:16603527 doi:87/5/1247
- ↑ Marzi A, Akhavan A, Simmons G, Gramberg T, Hofmann H, Bates P, Lingappa VR, Pohlmann S. The signal peptide of the ebolavirus glycoprotein influences interaction with the cellular lectins DC-SIGN and DC-SIGNR. J Virol. 2006 Jul;80(13):6305-17. PMID:16775318 doi:10.1128/JVI.02545-05
- ↑ Saeed MF, Kolokoltsov AA, Albrecht T, Davey RA. Cellular entry of ebola virus involves uptake by a macropinocytosis-like mechanism and subsequent trafficking through early and late endosomes. PLoS Pathog. 2010 Sep 16;6(9):e1001110. doi: 10.1371/journal.ppat.1001110. PMID:20862315 doi:10.1371/journal.ppat.1001110
- ↑ Bhattacharyya S, Warfield KL, Ruthel G, Bavari S, Aman MJ, Hope TJ. Ebola virus uses clathrin-mediated endocytosis as an entry pathway. Virology. 2010 May 25;401(1):18-28. doi: 10.1016/j.virol.2010.02.015. Epub 2010, Mar 3. PMID:20202662 doi:10.1016/j.virol.2010.02.015
- ↑ Yang ZY, Duckers HJ, Sullivan NJ, Sanchez A, Nabel EG, Nabel GJ. Identification of the Ebola virus glycoprotein as the main viral determinant of vascular cell cytotoxicity and injury. Nat Med. 2000 Aug;6(8):886-9. PMID:10932225 doi:10.1038/78645
- ↑ Alvarez CP, Lasala F, Carrillo J, Muniz O, Corbi AL, Delgado R. C-type lectins DC-SIGN and L-SIGN mediate cellular entry by Ebola virus in cis and in trans. J Virol. 2002 Jul;76(13):6841-4. PMID:12050398
- ↑ Wahl-Jensen VM, Afanasieva TA, Seebach J, Stroher U, Feldmann H, Schnittler HJ. Effects of Ebola virus glycoproteins on endothelial cell activation and barrier function. J Virol. 2005 Aug;79(16):10442-50. PMID:16051836 doi:79/16/10442
- ↑ Wahl-Jensen V, Kurz SK, Hazelton PR, Schnittler HJ, Stroher U, Burton DR, Feldmann H. Role of Ebola virus secreted glycoproteins and virus-like particles in activation of human macrophages. J Virol. 2005 Feb;79(4):2413-9. PMID:15681442 doi:79/4/2413
- ↑ Alazard-Dany N, Volchkova V, Reynard O, Carbonnelle C, Dolnik O, Ottmann M, Khromykh A, Volchkov VE. Ebola virus glycoprotein GP is not cytotoxic when expressed constitutively at a moderate level. J Gen Virol. 2006 May;87(Pt 5):1247-57. PMID:16603527 doi:87/5/1247
- ↑ Marzi A, Akhavan A, Simmons G, Gramberg T, Hofmann H, Bates P, Lingappa VR, Pohlmann S. The signal peptide of the ebolavirus glycoprotein influences interaction with the cellular lectins DC-SIGN and DC-SIGNR. J Virol. 2006 Jul;80(13):6305-17. PMID:16775318 doi:10.1128/JVI.02545-05
- ↑ Saeed MF, Kolokoltsov AA, Albrecht T, Davey RA. Cellular entry of ebola virus involves uptake by a macropinocytosis-like mechanism and subsequent trafficking through early and late endosomes. PLoS Pathog. 2010 Sep 16;6(9):e1001110. doi: 10.1371/journal.ppat.1001110. PMID:20862315 doi:10.1371/journal.ppat.1001110
- ↑ Bhattacharyya S, Warfield KL, Ruthel G, Bavari S, Aman MJ, Hope TJ. Ebola virus uses clathrin-mediated endocytosis as an entry pathway. Virology. 2010 May 25;401(1):18-28. doi: 10.1016/j.virol.2010.02.015. Epub 2010, Mar 3. PMID:20202662 doi:10.1016/j.virol.2010.02.015
- ↑ Lee JE, Fusco ML, Hessell AJ, Oswald WB, Burton DR, Saphire EO. Structure of the Ebola virus glycoprotein bound to an antibody from a human survivor. Nature. 2008 Jul 10;454(7201):177-82. PMID:18615077 doi:10.1038/nature07082
|