This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2f4m
From Proteopedia
| Line 4: | Line 4: | ||
|PDB= 2f4m |SIZE=350|CAPTION= <scene name='initialview01'>2f4m</scene>, resolution 1.85Å | |PDB= 2f4m |SIZE=350|CAPTION= <scene name='initialview01'>2f4m</scene>, resolution 1.85Å | ||
|SITE= | |SITE= | ||
| - | |LIGAND= <scene name='pdbligand= | + | |LIGAND= <scene name='pdbligand=CL:CHLORIDE+ION'>CL</scene>, <scene name='pdbligand=ZN:ZINC+ION'>ZN</scene> |
| - | |ACTIVITY= [http://en.wikipedia.org/wiki/Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine_amidase Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine amidase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=3.5.1.52 3.5.1.52] | + | |ACTIVITY= <span class='plainlinks'>[http://en.wikipedia.org/wiki/Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine_amidase Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine amidase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=3.5.1.52 3.5.1.52] </span> |
|GENE= Rad23b, Mhr23b ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=10090 Mus musculus]) | |GENE= Rad23b, Mhr23b ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=10090 Mus musculus]) | ||
| + | |DOMAIN= | ||
| + | |RELATEDENTRY=[[2f4o|2F4O]] | ||
| + | |RESOURCES=<span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=2f4m FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2f4m OCA], [http://www.ebi.ac.uk/pdbsum/2f4m PDBsum], [http://www.rcsb.org/pdb/explore.do?structureId=2f4m RCSB]</span> | ||
}} | }} | ||
| Line 29: | Line 32: | ||
[[Category: Zhao, G.]] | [[Category: Zhao, G.]] | ||
[[Category: Zhou, X.]] | [[Category: Zhou, X.]] | ||
| - | [[Category: CL]] | ||
| - | [[Category: ZN]] | ||
[[Category: glycoprotein]] | [[Category: glycoprotein]] | ||
[[Category: nucleotide excision repair]] | [[Category: nucleotide excision repair]] | ||
| Line 37: | Line 38: | ||
[[Category: ubiquitin-dependent protein degradation]] | [[Category: ubiquitin-dependent protein degradation]] | ||
| - | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on | + | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Mon Mar 31 02:57:48 2008'' |
Revision as of 23:57, 30 March 2008
| |||||||
| , resolution 1.85Å | |||||||
|---|---|---|---|---|---|---|---|
| Ligands: | , | ||||||
| Gene: | Rad23b, Mhr23b (Mus musculus) | ||||||
| Activity: | Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine amidase, with EC number 3.5.1.52 | ||||||
| Related: | 2F4O
| ||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
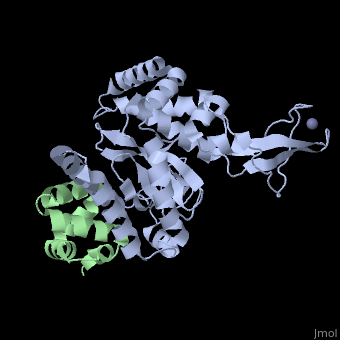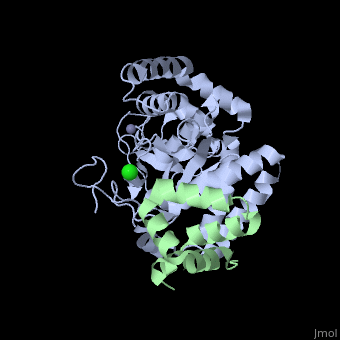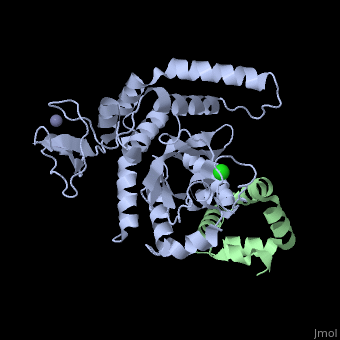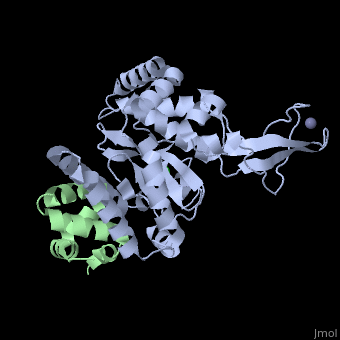
The Mouse PNGase-HR23 Complex Reveals a Complete Remodulation of the Protein-Protein Interface Compared to its Yeast Orthologs
Overview
Peptide N-glycanase removes N-linked oligosaccharides from misfolded glycoproteins as part of the endoplasmic reticulum-associated degradation pathway. This process involves the formation of a tight complex of peptide N-glycanase with Rad23 in yeast and the orthologous HR23 proteins in mammals. In addition to its function in endoplasmic reticulum-associated degradation, HR23 is also involved in DNA repair, where it plays an important role in damage recognition in complex with the xeroderma pigmentosum group C protein. To characterize the dual role of HR23, we have determined the high resolution crystal structure of the mouse peptide N-glycanase catalytic core in complex with the xeroderma pigmentosum group C binding domain from HR23B. Peptide N-glycanase features a large cleft between its catalytic cysteine protease core and zinc binding domain. Opposite the zinc binding domain is the HR23B-interacting region, and surprisingly, the complex interface is fundamentally different from the orthologous yeast peptide N-glycanase-Rad23 complex. Different regions on both proteins are involved in complex formation, revealing an amazing degree of divergence in the interaction between two highly homologous proteins. Furthermore, the mouse peptide N-glycanase-HR23B complex mimics the interaction between xeroderma pigmentosum group C and HR23B, thereby providing a first structural model of how the two proteins interact within the nucleotide excision repair cascade in higher eukaryotes. The different interaction interfaces of the xeroderma pigmentosum group C binding domains in yeast and mammals suggest a co-evolution of the endoplasmic reticulum-associated degradation and DNA repair pathways.
About this Structure
2F4M is a Protein complex structure of sequences from Mus musculus. Full crystallographic information is available from OCA.
Reference
Structure of the mouse peptide N-glycanase-HR23 complex suggests co-evolution of the endoplasmic reticulum-associated degradation and DNA repair pathways., Zhao G, Zhou X, Wang L, Li G, Kisker C, Lennarz WJ, Schindelin H, J Biol Chem. 2006 May 12;281(19):13751-61. Epub 2006 Feb 24. PMID:16500903
Page seeded by OCA on Mon Mar 31 02:57:48 2008
Categories: Mus musculus | Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine amidase | Protein complex | Kisker, C. | Lennarz, W J. | Schindelin, H. | Wang, L. | Zhao, G. | Zhou, X. | Glycoprotein | Nucleotide excision repair | Peptide:n-glycanase | Transglutaminase | Ubiquitin-dependent protein degradation