This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 972
From Proteopedia
(Difference between revisions)
| Line 45: | Line 45: | ||
==Hypothetical role of bradykinin on IDE== | ==Hypothetical role of bradykinin on IDE== | ||
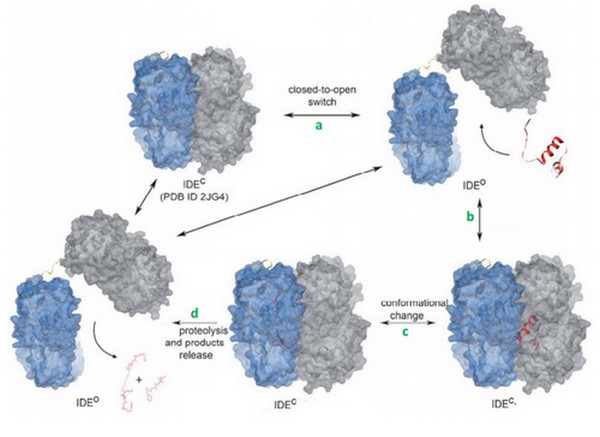
| - | Today, we can supposed that binding of bradykinin at the exosite stimulated the conformationnal change of IDE, from its open to its close state. | + | Today, we can supposed that binding of bradykinin at the exosite stimulated the conformationnal change of IDE<ref> doi: 10.1038/nature05143</ref>, from its open to its close state. |
We can also suggests that IDE binds <scene name='60/604491/Bradykinin/1'>2 bradykinins</scene> thanks to their small lenght. | We can also suggests that IDE binds <scene name='60/604491/Bradykinin/1'>2 bradykinins</scene> thanks to their small lenght. | ||
Binding of bradykinin or other short peptides to the exosite could play a regulatory role in substrate binding and cleavage by IDE<ref>doi: 10.1021/bi801192h</ref>. | Binding of bradykinin or other short peptides to the exosite could play a regulatory role in substrate binding and cleavage by IDE<ref>doi: 10.1021/bi801192h</ref>. | ||
Revision as of 21:54, 9 January 2015
| This Sandbox is Reserved from 15/11/2014, through 15/05/2015 for use in the course "Biomolecule" taught by Bruno Kieffer at the Strasbourg University. This reservation includes Sandbox Reserved 951 through Sandbox Reserved 975. |
To get started:
More help: Help:Editing |
| |||||||||||
References
- ↑ Song ES, Juliano MA, Juliano L, Hersh LB. Substrate activation of insulin-degrading enzyme (insulysin). A potential target for drug development. J Biol Chem. 2003 Dec 12;278(50):49789-94. Epub 2003 Oct 2. PMID:14527953 doi:http://dx.doi.org/10.1074/jbc.M308983200
- ↑ Im H, Manolopoulou M, Malito E, Shen Y, Zhao J, Neant-Fery M, Sun CY, Meredith SC, Sisodia SS, Leissring MA, Tang WJ. Structure of substrate-free human insulin-degrading enzyme (IDE) and biophysical analysis of ATP-induced conformational switch of IDE. J Biol Chem. 2007 Aug 31;282(35):25453-63. Epub 2007 Jul 5. PMID:17613531 doi:10.1074/jbc.M701590200
- ↑ Shen Y, Joachimiak A, Rosner MR, Tang WJ. Structures of human insulin-degrading enzyme reveal a new substrate recognition mechanism. Nature. 2006 Oct 19;443(7113):870-4. Epub 2006 Oct 11. PMID:17051221 doi:10.1038/nature05143
- ↑ Malito E, Ralat LA, Manolopoulou M, Tsay JL, Wadlington NL, Tang WJ. Molecular Bases for the Recognition of Short Peptide Substrates and Cysteine-Directed Modifications of Human Insulin-Degrading Enzyme. Biochemistry. 2008 Nov 6. PMID:18986166 doi:10.1021/bi801192h