This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Z-DNA
From Proteopedia
| Line 2: | Line 2: | ||
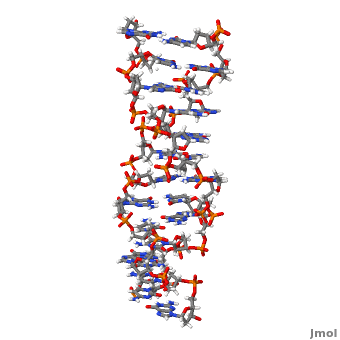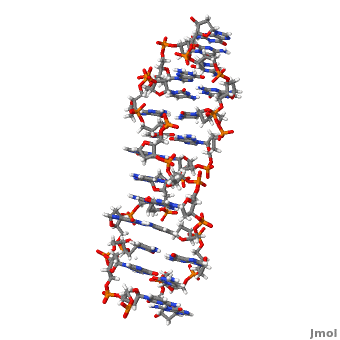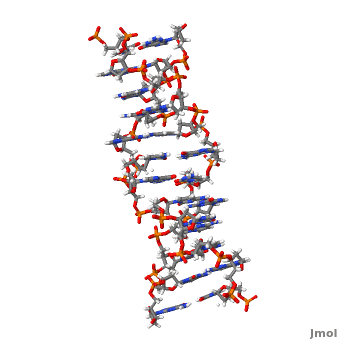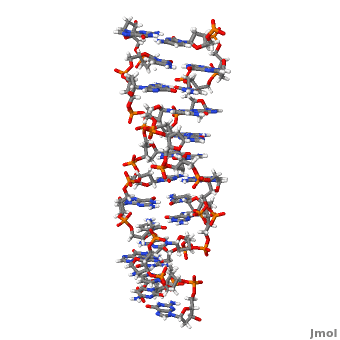
'''Z-DNA''' <scene name='Z-DNA/Z-dna_new/1'>(default scene)</scene> is a form of DNA that has a different structure from the more common <scene name='Sandbox_Z-DNA/Bdna/3'>B-DNA</scene> form.It is a left-handed double helix wherein the sugar-phosphate backbone has a zigzag pattern due to the alternate stacking of bases in [http://proteopedia.org/wiki/index.php/Syn_and_anti_nucleosides anti-conformation and syn conformation]. In Z-DNA only a minor groove is present and the major groove is absent. The residues that allow sequence-specific recognition of Z-DNA are present on the convex outer surface.<ref name = 'Rich'> PMID:12838348</ref> This DNA form is thought to play a role in the regulation of gene expression, DNA processing events and/or genetic instability.<ref name = 'Wang'>PMID:17485386</ref> | '''Z-DNA''' <scene name='Z-DNA/Z-dna_new/1'>(default scene)</scene> is a form of DNA that has a different structure from the more common <scene name='Sandbox_Z-DNA/Bdna/3'>B-DNA</scene> form.It is a left-handed double helix wherein the sugar-phosphate backbone has a zigzag pattern due to the alternate stacking of bases in [http://proteopedia.org/wiki/index.php/Syn_and_anti_nucleosides anti-conformation and syn conformation]. In Z-DNA only a minor groove is present and the major groove is absent. The residues that allow sequence-specific recognition of Z-DNA are present on the convex outer surface.<ref name = 'Rich'> PMID:12838348</ref> This DNA form is thought to play a role in the regulation of gene expression, DNA processing events and/or genetic instability.<ref name = 'Wang'>PMID:17485386</ref> | ||
| + | |||
| + | See also [[Z-DNA model tour]]. | ||
== Structure == | == Structure == | ||
| - | Z-DNA (<scene name='Sandbox_Z-DNA/B-z/7'>B-Z DNA junction</scene>, PDB entry [[2acj]]) can form '' | + | Z-DNA (<scene name='Sandbox_Z-DNA/B-z/7'>B-Z DNA junction</scene>, PDB entry [[2acj]]) can form ''in vitro'' from B-DNA by raising negative super helical stress or under low salt conditions when deoxycytosine is 5-methylated. The formation of Z-DNA ''in vivo'' is an energy requiring process. It forms behind a RNA polymerase moving through a DNA double helix during transcription and is subsequently stabilized due to the generation of negative supercoils. Z-DNA is the first single crystal X-ray structure of a DNA fragment. It was crystallized as a self complementary DNA hexamer d(CG)<sub>3</sub> by Andrew Wang, Alexander Rich and their co-workers at MIT in 1979. <ref name = 'Rich'>PMID:12838348</ref><ref name ='Wang'>PMID:17485386</ref> |
Whenever B-DNA transforms into Z-DNA two <scene name='Sandbox_Z-DNA/B-zjunction/7'>B-Z junctions</scene> form. The crystal structure of these junctions revealed<scene name='Sandbox_Z-DNA/Extruded/12'> two extruded bases</scene>, <scene name='Z-DNA/Extruded/2'>adenine</scene> and <scene name='Z-DNA/Extruded/3'>thymine</scene> at the junction. A crucial finding from this structure is that a right handed DNA can transform to a left handed DNA or vice versa by the disruption and extrusion of a base pair. It has also been suggested that the extruded base pairs at B-Z DNA junction may be sites for DNA modification.<ref>PMID:16237447</ref> | Whenever B-DNA transforms into Z-DNA two <scene name='Sandbox_Z-DNA/B-zjunction/7'>B-Z junctions</scene> form. The crystal structure of these junctions revealed<scene name='Sandbox_Z-DNA/Extruded/12'> two extruded bases</scene>, <scene name='Z-DNA/Extruded/2'>adenine</scene> and <scene name='Z-DNA/Extruded/3'>thymine</scene> at the junction. A crucial finding from this structure is that a right handed DNA can transform to a left handed DNA or vice versa by the disruption and extrusion of a base pair. It has also been suggested that the extruded base pairs at B-Z DNA junction may be sites for DNA modification.<ref>PMID:16237447</ref> | ||
Revision as of 21:13, 11 January 2017
| |||||||||||
Movie Depicting ADAR1 binding to Z-DNA
Comparison of the three helices and helical parameters of DNA
Sources[7]
|
|
|
| Parameter | A-DNA | B-DNA | Z-DNA |
|---|---|---|---|
| Helix sense | right-handed | right-handed | left-handed |
| Residues per turn | 11 | 10.5 | 12 |
| Axial rise [Å] | 2.55 | 3.4 | 3.7 |
| Helix pitch(°) | 28 | 34 | 45 |
| Base pair tilt(°) | 20 | −6 | 7 |
| Rotation per residue (°) | 33 | 36 | -30 |
| Diameter of helix [Å] | 23 | 20 | 18 |
| Glycosidic bond configuration dA,dT,dC dG | anti anti | anti anti | anti syn |
| Sugar pucker dA,dT,dC dG | C3'-endo C3'-endo | C2'-endo C2'-endo | C2'-endo C3'-endo |
| Intrastrand phosphate-phosphate distance [Å] dA,dT,dC dG | 5.9 5.9 | 7.0 7.0 | 7.0 5.9 |
| Sources:[8][9][10] | |||
3D structures of Z-DNA
Updated on 11-January-2017
1woe, 3qba – ZDNA hexamer CGCGCG
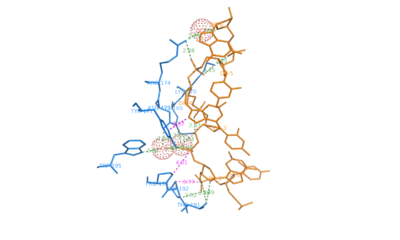
1qbj – ZDNA hexamer CGCGCG + hADAR1 Zα domain - human
2heo - ZDNA hexamer CGCGCG + Z-DNA-binding protein Zα domain – mouse
2eyi - ZDNA hexamer CGCGCG + Z-DNA-binding protein Zβ domain
1j75 - ZDNA hexamer CGCGCG + DLM-1 Zα domain
Additional Resources
For additional information, see: Nucleic Acids
References
- ↑ 1.0 1.1 1.2 Rich A, Zhang S. Timeline: Z-DNA: the long road to biological function. Nat Rev Genet. 2003 Jul;4(7):566-72. PMID:12838348 doi:10.1038/nrg1115
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 Wang G, Vasquez KM. Z-DNA, an active element in the genome. Front Biosci. 2007 May 1;12:4424-38. PMID:17485386
- ↑ Ha SC, Lowenhaupt K, Rich A, Kim YG, Kim KK. Crystal structure of a junction between B-DNA and Z-DNA reveals two extruded bases. Nature. 2005 Oct 20;437(7062):1183-6. PMID:16237447 doi:10.1038/nature04088
- ↑ Schwartz T, Rould MA, Lowenhaupt K, Herbert A, Rich A. Crystal structure of the Zalpha domain of the human editing enzyme ADAR1 bound to left-handed Z-DNA. Science. 1999 Jun 11;284(5421):1841-5. PMID:10364558
- ↑ Kim YG, Lowenhaupt K, Oh DB, Kim KK, Rich A. Evidence that vaccinia virulence factor E3L binds to Z-DNA in vivo: Implications for development of a therapy for poxvirus infection. Proc Natl Acad Sci U S A. 2004 Feb 10;101(6):1514-8. Epub 2004 Feb 2. PMID:14757814 doi:10.1073/pnas.0308260100
- ↑ Drozdzal P, Gilski M, Kierzek R, Lomozik L, Jaskolski M. High-resolution crystal structure of Z-DNA in complex with Cr cations. J Biol Inorg Chem. 2015 Feb 17. PMID:25687556 doi:http://dx.doi.org/10.1007/s00775-015-1247-5
- ↑ http://203.129.231.23/indira/nacc/
- ↑ Rich A, Nordheim A, Wang AH. The chemistry and biology of left-handed Z-DNA. Annu Rev Biochem. 1984;53:791-846. PMID:6383204 doi:http://dx.doi.org/10.1146/annurev.bi.53.070184.004043
- ↑ Wang AH, Quigley GJ, Kolpak FJ, Crawford JL, van Boom JH, van der Marel G, Rich A. Molecular structure of a left-handed double helical DNA fragment at atomic resolution. Nature. 1979 Dec 13;282(5740):680-6. PMID:514347
- ↑ Sinden, Richard R (1994-01-15). DNA structure and function (1st ed.). Academic Press. pp. 398. ISBN 0-12-645750-6.
Proteopedia Page Contributors and Editors (what is this?)
Adithya Sagar, Michal Harel, Joel L. Sussman, Alexander Berchansky, David Canner, Donald Voet, Jaime Prilusky, Karl Oberholser, Eran Hodis