Introduction
Isocitrate lyase is a lyase found in the proteome of multiple bacteria that oxidizes the hydroxl group of isocitrate and cleaves the substrate in two forming glyoxylate and succinate. Isocitrate lyase is a tetramer that is composed primarily of alpha helices and beta sheets with a unqiue structural phenomenon called "helix swapping".
Protein Structure
Crystal Structure
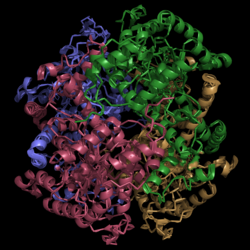
Figure 2. Structure of Isocitrate Lyase. Quaternary structure is comprised of four subunits forming an alpha/beta barrel.
Isocitrate lyase (PDB Code 1F8I) is a tetramer with 222 symmetry. Each subunit is composed of 14 alpha helices and 14 beta sheets. A unique structural feature of this enzyme is a phenomenon called "".
Helix swapping is observed between two monomers to form stable dimers. The 11th and 12th helices of each monomer exchange three dimensional placement with the respective helices of the opposite monomer. Due to the 222 symmetry observed, only two dimers form than combine to form the observed tetramer. As a result of this structure, 18% of the surface of each monomer is buried within the protein.
Active Site Residues
Inhibitors
Mechanism of Action
Image:TCA Cycle.png Figure 1. Citric Acid Cycle with Glyoxylate Shunt Pathway. In several bacterial species, there is a carbon conserving gloxylate shunt pathway that converts isocitrate to malate in two steps instead of the usual five steps.
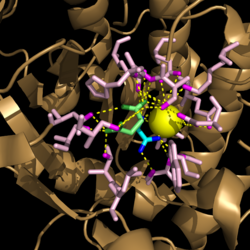
Figure 3. Active site residues hydrogen bound to a cofactor and the products of the catalyzed isocitrate reaction. Glyoxylate is shown in blue, succinate is shown in green, and the Mg
2+ cofactor is shown in yellow.
Disease Association
Clinical Implications
Mycobacterium tuberculosis is a respiratory infection that causes numerous fatalities throughout the world. It lives in organisms and feeds off of host cells, which indicate a variety of lipases exist within M. tuberculosis. Current drugs that are on the market now target a small number of bacterial processes like cell wall formation and chromosomal replication. Although several antibiotics exist, all of them target these same mechanisms of inhibition. These commonalities have led to the prevalence of different multi-drug resistant (MDR) tuberculosis strains. Due to the high level of resistance, finding a lasting treatment for MDR TB infections has become very problematic. Studies into new mechanisms of inhibition will be crucial to prevent widespread outbreaks.
Isocitrate lyase plays a key role in survival of M. tuberculosis by sustaining intracellular infections in inflammatory respiratory macrophages. Used in the citric acid cycle, isocitrate lyase is the first enzyme catalyzing the carbon conserving glyoxylate pathway. This glyoxylate pathway has not been observed in mammals and thus presents a unique drug target to solely attack TB infections.


