This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1058
From Proteopedia
(Difference between revisions)
| Line 13: | Line 13: | ||
===Active Site Residues=== | ===Active Site Residues=== | ||
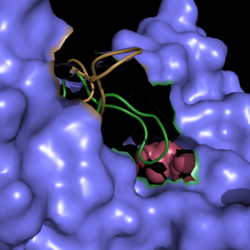
| - | [[Image:Active Loop Shift.png|250 px|center|thumb|'''Figure 1. Active Site Loop Shift.''' Binding of the ligand to the enzyme results in a conformational shift that facilitates the breakdown of isocitrate. The active site loop unbound is shown in | + | [[Image:Active Loop Shift.png|250 px|center|thumb|'''Figure 1. Active Site Loop Shift.''' Binding of the ligand to the enzyme results in a conformational shift that facilitates the breakdown of isocitrate. The active site loop unbound is shown in wheat and the active site loop bound is shown in green.]] |
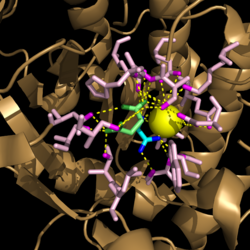
[[Image:Active_Site_Hydrogen_Bonding.png|250 px|left|thumb|'''Figure 3. Active site residues hydrogen bound to a cofactor and the products of the catalyzed isocitrate reaction.''' Glyoxylate is shown in blue, succinate is shown in green, and the Mg<sup>2+</sup> cofactor is shown in yellow.]] | [[Image:Active_Site_Hydrogen_Bonding.png|250 px|left|thumb|'''Figure 3. Active site residues hydrogen bound to a cofactor and the products of the catalyzed isocitrate reaction.''' Glyoxylate is shown in blue, succinate is shown in green, and the Mg<sup>2+</sup> cofactor is shown in yellow.]] | ||
Revision as of 19:36, 7 April 2015
Isocitrate Lyase from Mycobacterium tuberculosis
| |||||||||||