This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1072
From Proteopedia
(Difference between revisions)
| Line 8: | Line 8: | ||
===Catalase Peroxidases=== | ===Catalase Peroxidases=== | ||
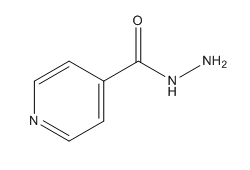
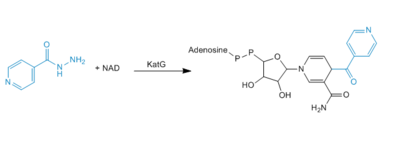
Catalase-peroxidases are enzymes that degrade hydrogen peroxide. Catalase converts two equivalents of hydrogen peroxide into water and oxygen via a two-step reaction cycle in which H<sub>2</sub>0<sub>2</sub> alternately oxidizes and reduces the heme iron at the active site. Within peroxidases, oxidation of heme iron involves a H<sub>2</sub>0<sub>2</sub> molecules, similar to that in the catalase-catalyzed reaction. Reduction of the heme iron, however, involves hydrogen donors such as NADH, not a second H<sub>2</sub>0<sub>2</sub> molcule (3). Catalase-Peroxidases that have been characterized are either homodimers or homotetramers and contain a single heme ''b'' cofactor at the active site. Usually, the primary struture of the subunit can be divided into two halves that have a high level of sequence similarity, most likely due to a gene duplication event. | Catalase-peroxidases are enzymes that degrade hydrogen peroxide. Catalase converts two equivalents of hydrogen peroxide into water and oxygen via a two-step reaction cycle in which H<sub>2</sub>0<sub>2</sub> alternately oxidizes and reduces the heme iron at the active site. Within peroxidases, oxidation of heme iron involves a H<sub>2</sub>0<sub>2</sub> molecules, similar to that in the catalase-catalyzed reaction. Reduction of the heme iron, however, involves hydrogen donors such as NADH, not a second H<sub>2</sub>0<sub>2</sub> molcule (3). Catalase-Peroxidases that have been characterized are either homodimers or homotetramers and contain a single heme ''b'' cofactor at the active site. Usually, the primary struture of the subunit can be divided into two halves that have a high level of sequence similarity, most likely due to a gene duplication event. | ||
| + | |||
==Structure== | ==Structure== | ||
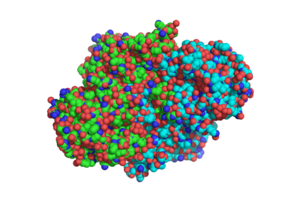
[[Image:DotSTRUCTURE.png|300 px|left|thumb|Overall structure of [http://www.rcsb.org/pdb/explore/explore.do?structureId=1sJ2 1SJ2], showing that the structure is a [http://en.wikipedia.org/wiki/Protein_dimer homodimer].]] | [[Image:DotSTRUCTURE.png|300 px|left|thumb|Overall structure of [http://www.rcsb.org/pdb/explore/explore.do?structureId=1sJ2 1SJ2], showing that the structure is a [http://en.wikipedia.org/wiki/Protein_dimer homodimer].]] | ||
| - | ==Structure== | ||
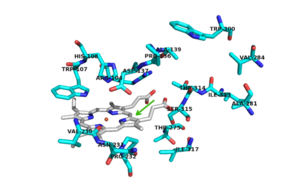
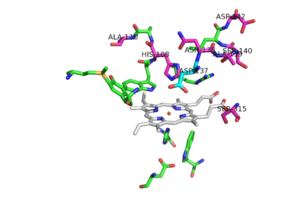
The two [http://en.wikipedia.org/wiki/Protein_domain domains] of each monomer are primarily [http://en.wikipedia.org/wiki/Alpha_helix alpha helical] and have similar foldings. The similar foldings suggests that the monomer results from a [http://en.wikipedia.org/wiki/Gene_duplication gene duplication] event; however, the C-terminal domain does not contain the [http://en.wikipedia.org/wiki/Heme_B heme ''b''] prosthetic group, while the <scene name='69/694238/N_terminus/1'>N terminal</scene> does. The [http://en.wikipedia.org/wiki/Active_site active site] is therefore located within the N-terminal domain. The two monomers interact through an interlocking hook formed by the N-terminal domains that stabilizes the formation of the dimer (1). | The two [http://en.wikipedia.org/wiki/Protein_domain domains] of each monomer are primarily [http://en.wikipedia.org/wiki/Alpha_helix alpha helical] and have similar foldings. The similar foldings suggests that the monomer results from a [http://en.wikipedia.org/wiki/Gene_duplication gene duplication] event; however, the C-terminal domain does not contain the [http://en.wikipedia.org/wiki/Heme_B heme ''b''] prosthetic group, while the <scene name='69/694238/N_terminus/1'>N terminal</scene> does. The [http://en.wikipedia.org/wiki/Active_site active site] is therefore located within the N-terminal domain. The two monomers interact through an interlocking hook formed by the N-terminal domains that stabilizes the formation of the dimer (1). | ||
Revision as of 13:15, 14 April 2015
| This Sandbox is Reserved from 02/09/2015, through 05/31/2016 for use in the course "CH462: Biochemistry 2" taught by Geoffrey C. Hoops at the Butler University. This reservation includes Sandbox Reserved 1051 through Sandbox Reserved 1080. |
To get started:
More help: Help:Editing |
| |||||||||||