This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1056
From Proteopedia
(Difference between revisions)
| Line 2: | Line 2: | ||
<StructureSection load='1F8I' size='340' side='right' caption='Isocitrate Lyase from ''Mycobacterium tuberculosis''' scene='Isocitrate Lyase complex with glyoxylate and succinate ligands bound'> | <StructureSection load='1F8I' size='340' side='right' caption='Isocitrate Lyase from ''Mycobacterium tuberculosis''' scene='Isocitrate Lyase complex with glyoxylate and succinate ligands bound'> | ||
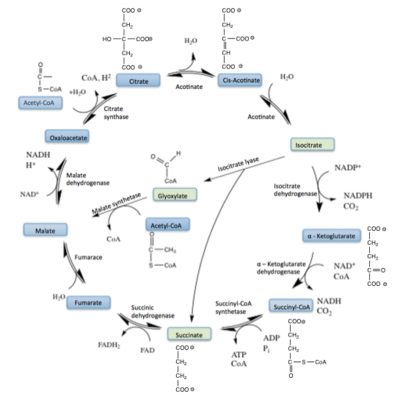
| - | [[Image: | + | [[Image:Glyox_Shunt.png|400 px|right|thumb|Figure 1: ICL mediated glyoxylate shunt pathway of the Citric Acid Cycle]] |
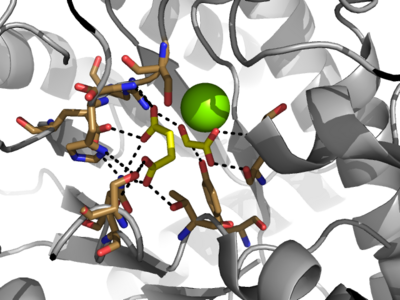
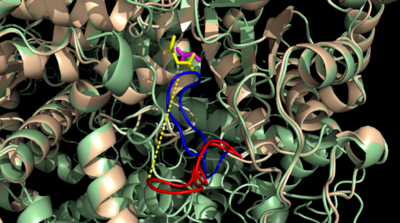
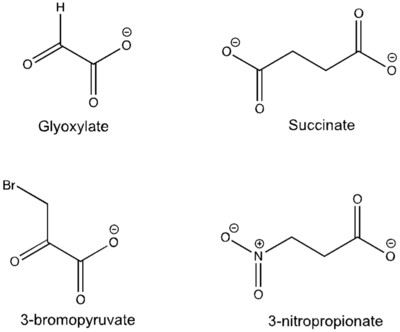
[http://en.wikipedia.org/wiki/Isocitrate_lyase Isocitrate Lyase] (ICL) is a metabolic enzyme that converts the metabolite isocitrate into glyoxylate and succinate. ICL is a homotetramer with each monomer being composed of 14 alpha helices, 14 beta sheets, and a magnesium ion cofactor. ICL has shown clinical relevance in the disease state [http://en.wikipedia.org/wiki/Tuberculosis Tuberculosis] where it is responsible for the persistence of ''Mycobacterium tuberculosis'' during the chronic stage of infection<ref name="genes">PMID: 18054522</ref> This survival strategy mediated by ICL is characterized by a metabolic shortcut within the [http://en.wikipedia.org/wiki/Citric_acid_cycle Citric Acid Cycle]. ICL creates this shunt pathway by converting isocitrate to succinate and glyoxylate, diverting acetyl-CoA from the beta-oxidation of fatty acids<ref name="ICL">PMID:10932251</ref><ref name="ICL2">PMID: 2696959</ref>. | [http://en.wikipedia.org/wiki/Isocitrate_lyase Isocitrate Lyase] (ICL) is a metabolic enzyme that converts the metabolite isocitrate into glyoxylate and succinate. ICL is a homotetramer with each monomer being composed of 14 alpha helices, 14 beta sheets, and a magnesium ion cofactor. ICL has shown clinical relevance in the disease state [http://en.wikipedia.org/wiki/Tuberculosis Tuberculosis] where it is responsible for the persistence of ''Mycobacterium tuberculosis'' during the chronic stage of infection<ref name="genes">PMID: 18054522</ref> This survival strategy mediated by ICL is characterized by a metabolic shortcut within the [http://en.wikipedia.org/wiki/Citric_acid_cycle Citric Acid Cycle]. ICL creates this shunt pathway by converting isocitrate to succinate and glyoxylate, diverting acetyl-CoA from the beta-oxidation of fatty acids<ref name="ICL">PMID:10932251</ref><ref name="ICL2">PMID: 2696959</ref>. | ||
Revision as of 18:20, 19 April 2015
Isocitrate Lyase from Mycobacterium Tuberculosis
| |||||||||||
3D Structures of Isocitrate Lyase
Updated on 19-April-2015
- ICL from other bacteria
References
- ↑ Srivastava V, Jain A, Srivastava BS, Srivastava R. Selection of genes of Mycobacterium tuberculosis upregulated during residence in lungs of infected mice. Tuberculosis (Edinb). 2008 May;88(3):171-7. Epub 2007 Dec 3. PMID:18054522 doi:http://dx.doi.org/10.1016/j.tube.2007.10.002
- ↑ 2.00 2.01 2.02 2.03 2.04 2.05 2.06 2.07 2.08 2.09 2.10 Sharma V, Sharma S, Hoener zu Bentrup K, McKinney JD, Russell DG, Jacobs WR Jr, Sacchettini JC. Structure of isocitrate lyase, a persistence factor of Mycobacterium tuberculosis. Nat Struct Biol. 2000 Aug;7(8):663-8. PMID:10932251 doi:10.1038/77964
- ↑ 3.0 3.1 3.2 Beeching JR. High sequence conservation between isocitrate lyase from Escherichia coli and Ricinus communis. Protein Seq Data Anal. 1989 Dec;2(6):463-6. PMID:2696959
- ↑ 4.0 4.1 4.2 4.3 Masamune et al. Bio-Claisen condensation catalyzed by thiolase from Zoogloea ramigera. Active site cysteine residues. "Journal of the American Chemical Society" 111: 1879-1881 (1989). DOI: 10.1021/ja00187a053
- ↑ Connely, M. L. Solvent-accessible surfaces of proteins and nucleic acids "Science" 221:709-713 (1983). DOI: 10.1126/science.6879170