This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Regulator of G protein signaling
From Proteopedia
(Difference between revisions)
| Line 34: | Line 34: | ||
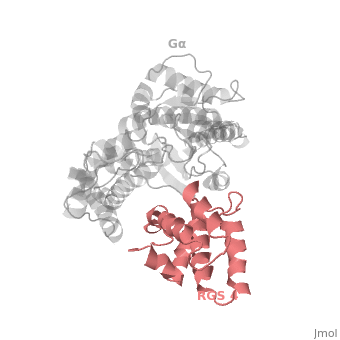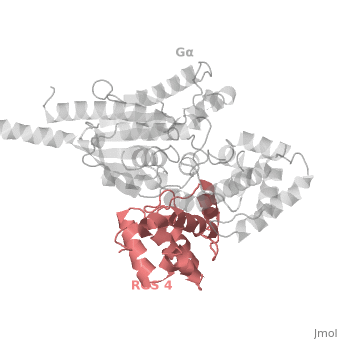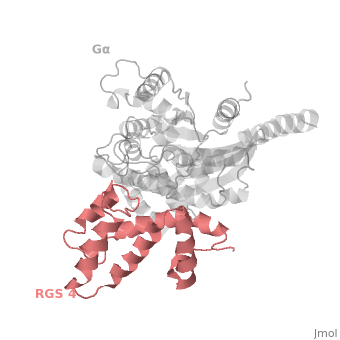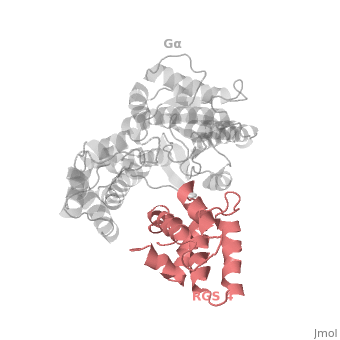
The <scene name='70/701447/Rgs4_monomer/4'>Monomer structure of RGS4</scene> in green cartoon diagram corresponds to an array of nine α-helices that fold into two small subdomains. The terminal subdomain contains the N and C termini of the box and is formed by α1, α2, α3, α8, and α9. Helices α1 and α9 lie in antiparallel orientation, juxtaposing the N and C termini of the box. The larger bundle subdomain, formed by α4, α5, α6, and α7, is a classic right-handed, antiparallel four-helix bundle. Both subdomains are required for GAP activity. | The <scene name='70/701447/Rgs4_monomer/4'>Monomer structure of RGS4</scene> in green cartoon diagram corresponds to an array of nine α-helices that fold into two small subdomains. The terminal subdomain contains the N and C termini of the box and is formed by α1, α2, α3, α8, and α9. Helices α1 and α9 lie in antiparallel orientation, juxtaposing the N and C termini of the box. The larger bundle subdomain, formed by α4, α5, α6, and α7, is a classic right-handed, antiparallel four-helix bundle. Both subdomains are required for GAP activity. | ||
| - | Gα<sub>i1</sub> subunits adopt a conserved fold composed of <scene name='70/701447/All-helical-domain/6'>α helical domain</scene> , a helical domain of six α helices shown as blue cartoon and a GTPase domain shown in gray cartoons.The GTPase domain hydrolyzes GTP and provides most of Gα's binding surfaces for Gβγ, receptors, effectors and RGS proteins.<scene name='70/701447/Gi-rgs4/19'>The GTPase domain</scene> contains three flexible regions designated switch-I presented as blue sticks, switch-II presented as magenta sticks and switch-III presented as green sticks that change conformation in response to GTP binding and hydrolysis. The three switch regions of Gα<sub>i1</sub>: residues 176–184, 201–215, and 233–241, respectively . <ref>PMID: 9108480</ref> | + | Gα<sub>i1</sub> subunits adopt a conserved fold composed of <scene name='70/701447/All-helical-domain/6'>α helical domain</scene> , a helical domain of six α helices shown as blue cartoon and a GTPase domain shown in gray cartoons.The GTPase domain hydrolyzes GTP and provides most of Gα's binding surfaces for Gβγ, receptors, effectors and RGS proteins. <scene name='70/701447/Gi-rgs4/19'>The GTPase domain</scene> contains three flexible regions designated switch-I presented as blue sticks, switch-II presented as magenta sticks and switch-III presented as green sticks that change conformation in response to GTP binding and hydrolysis. The three switch regions of Gα<sub>i1</sub>: residues 176–184, 201–215, and 233–241, respectively . <ref>PMID: 9108480</ref> |
</StructureSection> | </StructureSection> | ||
Revision as of 20:58, 18 May 2015
Regulator of G protein signaling (RGS) interactions with G proteins – RGS4-Gαi as a model structure.
| |||||||||||
References
- ↑ Kosloff M, Travis AM, Bosch DE, Siderovski DP, Arshavsky VY. Integrating energy calculations with functional assays to decipher the specificity of G protein-RGS protein interactions. Nat Struct Mol Biol. 2011 Jun 19;18(7):846-53. doi: 10.1038/nsmb.2068. PMID:21685921 doi:http://dx.doi.org/10.1038/nsmb.2068
- ↑ Milligan G, Kostenis E. Heterotrimeric G-proteins: a short history. Br J Pharmacol. 2006 Jan;147 Suppl 1:S46-55. PMID:16402120 doi:http://dx.doi.org/10.1038/sj.bjp.0706405
- ↑ Tesmer JJ, Berman DM, Gilman AG, Sprang SR. Structure of RGS4 bound to AlF4--activated G(i alpha1): stabilization of the transition state for GTP hydrolysis. Cell. 1997 Apr 18;89(2):251-61. PMID:9108480
Proteopedia Page Contributors and Editors (what is this?)
Ali Asli, Denise Salem, Michal Harel, Joel L. Sussman, Jaime Prilusky