This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Aminomutase
From Proteopedia
(Difference between revisions)
| Line 2: | Line 2: | ||
'''Aminomutases''' catalyze the exchange of hydrogen atom and an amine substituent on a neighboring carbon atom. | '''Aminomutases''' catalyze the exchange of hydrogen atom and an amine substituent on a neighboring carbon atom. | ||
| - | '''Phenylalanine aminomutase''' (PAM) catalyzes the reversible conversion of L-phenylalanine to ammonia and trans-cinnamic acid. Ala-Ser-Gly derived chromophore is a cofactor. | + | *'''Phenylalanine aminomutase''' (PAM) catalyzes the reversible conversion of L-phenylalanine to ammonia and trans-cinnamic acid. Ala-Ser-Gly derived chromophore is a cofactor. |
| - | '''Tyrosine-2,3-aminomutase''' (TAM) catalyzes the reversible conversion of L-tyrosine to 3-amino-3-(4-hydroxyphenyl)propanoate. Ala-Ser-Gly derived chromophore is a cofactor. | + | *'''Tyrosine-2,3-aminomutase''' (TAM) catalyzes the reversible conversion of L-tyrosine to 3-amino-3-(4-hydroxyphenyl)propanoate. Ala-Ser-Gly derived chromophore is a cofactor. |
| - | '''Ornithine-4,5-aminomutase''' (ORA) catalyzes the reversible conversion of D-ornithine to (2R,4S)-2,4-diaminopropanoate. ORA cofactors are pyridoxal phosphate (PLP), cobamide coenzyme and dithiothreitol. ORA contains subunits S (ORAS) and E (ORAE). The ORAS forms a complex with lysine 5,6-aminomutase. The ORAE contains a balamin-binding motif. | + | *'''Ornithine-4,5-aminomutase''' (ORA) catalyzes the reversible conversion of D-ornithine to (2R,4S)-2,4-diaminopropanoate. ORA cofactors are pyridoxal phosphate (PLP), cobamide coenzyme and dithiothreitol. ORA contains subunits S (ORAS) and E (ORAE). The ORAS forms a complex with lysine 5,6-aminomutase. The ORAE contains a balamin-binding motif. |
| - | '''Lysine-2,3-aminomutase''' (LAM) catalyzes the reversible conversion of lysine to beta-lysine. LAM is a S-adenosyl methionine (SAM) radical enzyme. LAM cofactors are PLP, Zn ion and FeS4 cluster. | + | *'''Lysine-2,3-aminomutase''' (LAM) catalyzes the reversible conversion of lysine to beta-lysine. LAM is a S-adenosyl methionine (SAM) radical enzyme. LAM cofactors are PLP, Zn ion and FeS4 cluster. |
| - | '''D-lysine-5,6-aminomutase''' (DLAM) catalyzes the reversible conversion of D-lysine to 2,5-diaminohexanoate. DLAM cofactor is cobamide | + | *'''D-lysine-5,6-aminomutase''' (DLAM) catalyzes the reversible conversion of D-lysine to 2,5-diaminohexanoate. DLAM cofactor is cobamide. |
| - | + | ||
| - | + | ||
| + | *'''Glutamate-1-semialdehyde 2,1-aminomutase''' (GSAM) catalyzes the reversible conversion of L-glutamate-1-semialdehyde to 5-aminovulinate. GSAM cofactor is PLP. | ||
== Function == | == Function == | ||
Revision as of 09:11, 2 December 2015
| |||||||||||
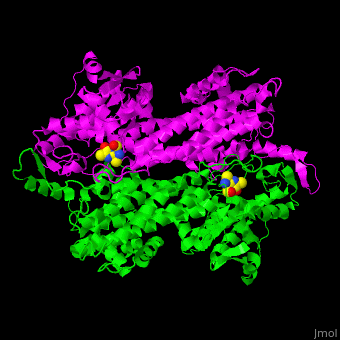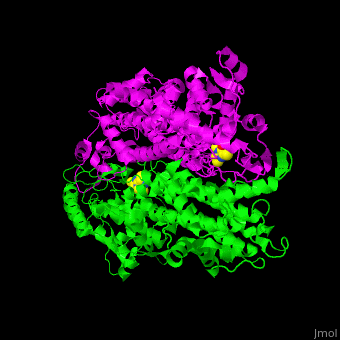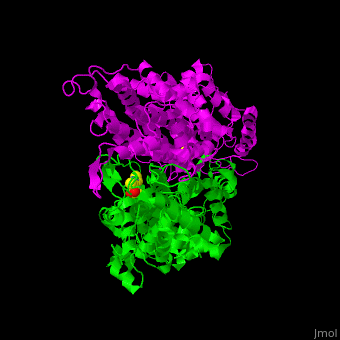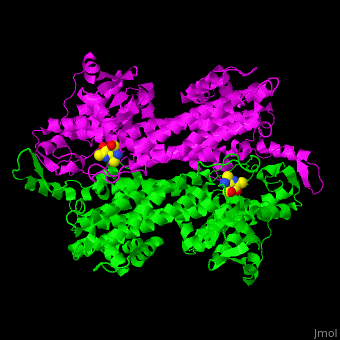
3D structures of aminomutase
Updated on 02-December-2015