This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
14-3-3 protein
From Proteopedia
(Difference between revisions)
| Line 15: | Line 15: | ||
== Structural highlights == | == Structural highlights == | ||
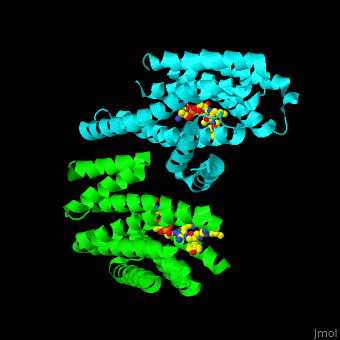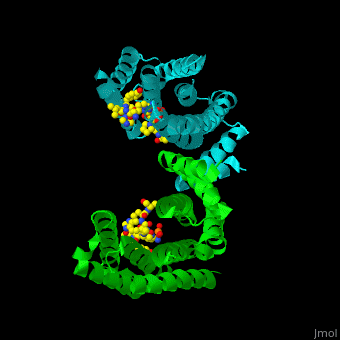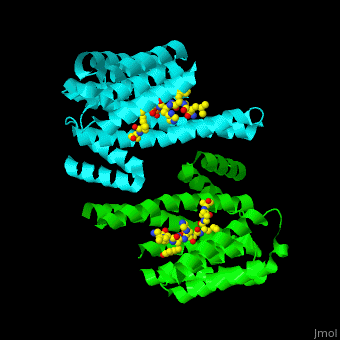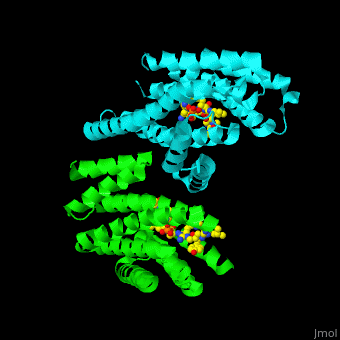
| - | PRS are homo- and heterodimers containing <scene name='59/590827/Cv/5'>9 antiparallel α-helices</scene>. <scene name='59/590827/Cv/12'>Three of the helices form the dimerization domain</scene> (<font color='red'><b>3 helices of chain A are in red</b></font> and <font color='magenta'><b>3 helices of chain B are in magenta</b></font>). <scene name='59/590827/Cv/ | + | PRS are homo- and heterodimers containing <scene name='59/590827/Cv/5'>9 antiparallel α-helices</scene>. <scene name='59/590827/Cv/12'>Three of the helices form the dimerization domain</scene> (<font color='red'><b>3 helices of chain A are in red</b></font> and <font color='magenta'><b>3 helices of chain B are in magenta</b></font>). <scene name='59/590827/Cv/13'>Five residues (in PRS-σ and PRS-ζ) are involved in ligand binding</scene>. |
</StructureSection> | </StructureSection> | ||
Revision as of 10:59, 13 December 2015
| |||||||||||
3D structures of 14-3-3 proteins (Updated on 13-December-2015)
References
- ↑ Benzinger A, Popowicz GM, Joy JK, Majumdar S, Holak TA, Hermeking H. The crystal structure of the non-liganded 14-3-3sigma protein: insights into determinants of isoform specific ligand binding and dimerization. Cell Res. 2005 Apr;15(4):219-27. PMID:15857576 doi:10.1038/sj.cr.7290290