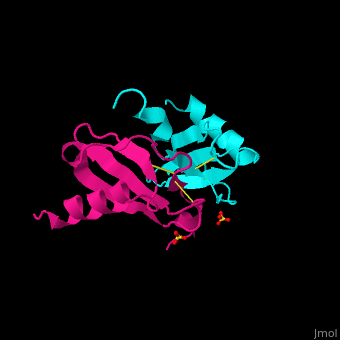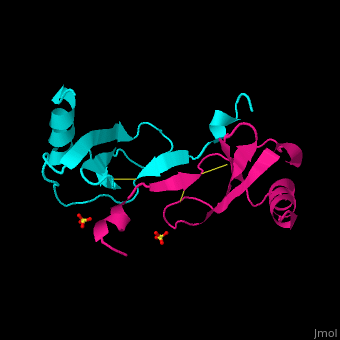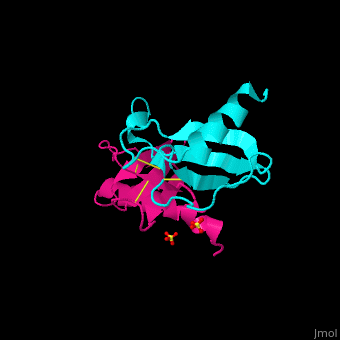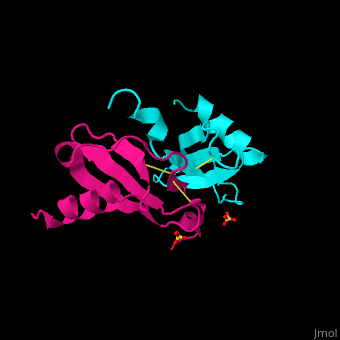This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Monocyte chemoattractant protein
From Proteopedia
| Line 14: | Line 14: | ||
It exists as a monomer or a dimer, eventhough the homodimer form is preferred. | It exists as a monomer or a dimer, eventhough the homodimer form is preferred. | ||
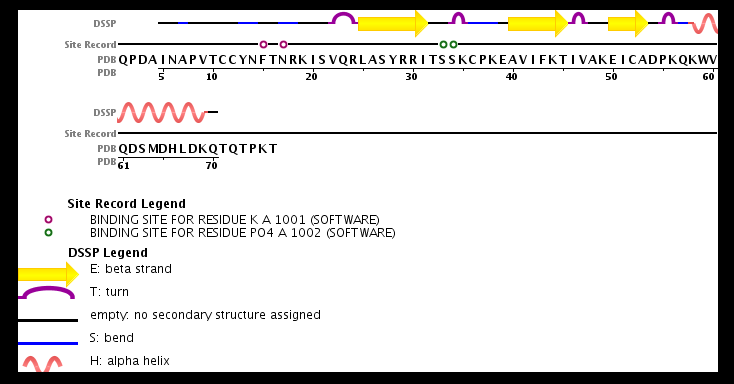
| - | + | The structure of the monomer is made of <scene name='72/721520/Ligand_binding_on_ccl2/2'>3 Beta sheets and 1 alpha helix</scene>. | |
| - | + | ||
| - | <scene name='72/721520/Ligand_binding_on_ccl2/2'>Beta sheets and alpha helix | + | |
== Ligands == | == Ligands == | ||
The known ligands for CCL2 are <scene name='72/721520/Ligand_binding_on_ccl2/1'>Potassium and PO4</scene>. | The known ligands for CCL2 are <scene name='72/721520/Ligand_binding_on_ccl2/1'>Potassium and PO4</scene>. | ||
Revision as of 12:00, 27 January 2016
| |||||||||||
Synthesis
The protein human CCL2 has been synthesized using a combination of solid phase peptide synthesis (SPPS) and native chemical ligation (NCL). The thioester-peptide segment was synthesized using the sulfonamide safety-catch linker and 9-fluorenylmethoxycarbonyl (Fmoc) SPPS, and pseudoproline dipeptides were used to facilitate the synthesis of both CCL2 fragments. After assembly of the full-length peptide chain by NCL, a glutathione redox buffer was used to fold and oxidize the CCL2 protein. CCL2 was crystallized and the structure was determined by X-ray diffraction at 1.9-A resolution. This is probably one of the first crystal structures of a protein prepared using the sulfonamide safety-catch linker and NCL.
3D structures of Monocyte chemoattractant protein
Updated on 27-January-2016
References
https://fr.wikipedia.org/wiki/CCL2 http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/pdbsum/GetPage.pl?pdbcode=1DOK http://www.uniprot.org/uniprot/P13500#interaction http://www.rcsb.org/pdb/explore/explore.do?structureId=3IFD
- ↑ Carr MW, Roth SJ, Luther E, Rose SS, Springer TA. Monocyte chemoattractant protein 1 acts as a T-lymphocyte chemoattractant. Proc Natl Acad Sci U S A. 1994 Apr 26;91(9):3652-6. PMID:8170963
- ↑ Ito Y, Ishiguro H, Kobayashi N, Hasumi H, Watanabe M, Yao M, Uemura H. Adipocyte-derived monocyte chemotactic protein-1 (MCP-1) promotes prostate cancer progression through the induction of MMP-2 activity. Prostate. 2015 Jul 1;75(10):1009-19. doi: 10.1002/pros.22972. Epub 2015 Apr 27. PMID:25917126 doi:http://dx.doi.org/10.1002/pros.22972
- ↑ Lubkowski J, Bujacz G, Boque L, Domaille PJ, Handel TM, Wlodawer A. The structure of MCP-1 in two crystal forms provides a rare example of variable quaternary interactions. Nat Struct Biol. 1997 Jan;4(1):64-9. PMID:8989326
Proteopedia Page Contributors and Editors (what is this?)
Coline Perrin, Alexane Caignard, Michal Harel, Alexander Berchansky, Joel L. Sussman