This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1136
From Proteopedia
| Line 1: | Line 1: | ||
{{Sandbox_Reserved_ESBS_2015}} | {{Sandbox_Reserved_ESBS_2015}} | ||
==Structure and Functional aspects of Sucrose Synthase from ''Arabidopsis thaliana''== | ==Structure and Functional aspects of Sucrose Synthase from ''Arabidopsis thaliana''== | ||
| - | Sucrose Synthase 1 (EC:2.4.1.13), also known as the Sucrose-UDP glucolsyltransferase 1 or simply Susy, is a reversible enzyme allowing the synthesis or the degradation of Sucrose in ''Arabidopsis thaliana''. It belongs to the Glycosyltransferase | + | Sucrose Synthase 1 (EC:2.4.1.13), also known as the Sucrose-UDP glucolsyltransferase 1 or simply Susy, is a reversible enzyme allowing the synthesis or the degradation of Sucrose in ''Arabidopsis thaliana''. It belongs to the Glycosyltransferase subfamily 4 (GT4). |
| - | <StructureSection scene=' | + | <StructureSection scene='' size='340' side='right' caption='X-ray crystal structures of AtSus1, as a complex with UDP-glucose at 2.8-Å resolution and as a complex with UDP and fructose at 2.85-Å resolution'> |
| + | |||
| + | 71/719877/Susy/3 | ||
== Function == | == Function == | ||
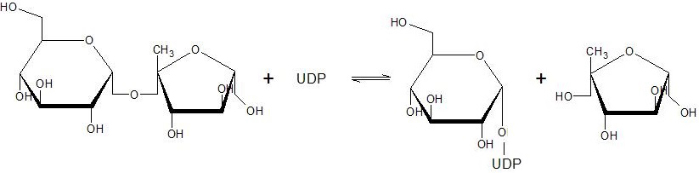
The Sucrose Synthase is able to catalyse the following reaction in both directions: | The Sucrose Synthase is able to catalyse the following reaction in both directions: | ||
| - | '''sucrose = glucose + fructose''' | ||
[[Image:Susy_reaction.jpg]] | [[Image:Susy_reaction.jpg]] | ||
| Line 27: | Line 28: | ||
• 1-127: N-Terminal regulatory domain involved in targeting <ref>Determination of structural requirements and probable regulatory effectors for membrane association of maize sucrose synthase 1. Hardin SC, Duncan KA, Huber SC. Plant Physiol. 2006</ref>. On this sequence, two serines can be phosphorylated, which enable a control of enzyme location <ref>Phosphorylation of sucrose synthase at serine 170: occurrence and possible role as a signal for proteolysis. Hardin SC, Tang GQ, Scholz A, Holtgraewe D, Winter H, Huber SC. Plant J. 2003</ref>. | • 1-127: N-Terminal regulatory domain involved in targeting <ref>Determination of structural requirements and probable regulatory effectors for membrane association of maize sucrose synthase 1. Hardin SC, Duncan KA, Huber SC. Plant Physiol. 2006</ref>. On this sequence, two serines can be phosphorylated, which enable a control of enzyme location <ref>Phosphorylation of sucrose synthase at serine 170: occurrence and possible role as a signal for proteolysis. Hardin SC, Tang GQ, Scholz A, Holtgraewe D, Winter H, Huber SC. Plant J. 2003</ref>. | ||
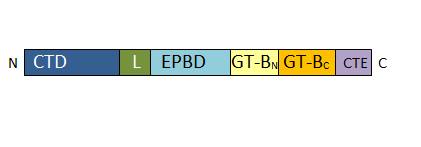
| - | • | + | • 1-127: CTD: Cellular targeting domain |
| - | • | + | • 157-276: EPBD: ENOD40 peptide-binding domain. This domain has a role in the regulation of the enzyme. |
| - | • 776-808 : C-terminal | + | • 277-776 : GT-B glycosyltransferase domain. It contains the catlytic site. |
| + | |||
| + | • 776-808 : C-terminal extension. The length of this domain is variable depending of the SUS isoform. | ||
| + | |||
| + | [[Image:Monomer_structure.jpg]] | ||
| + | |||
| + | == Structural highlights == | ||
Revision as of 11:53, 30 January 2016
| This Sandbox is Reserved from 15/12/2015, through 15/06/2016 for use in the course "Structural Biology" taught by Bruno Kieffer at the University of Strasbourg, ESBS. This reservation includes Sandbox Reserved 1120 through Sandbox Reserved 1159. |
To get started:
More help: Help:Editing |
Structure and Functional aspects of Sucrose Synthase from Arabidopsis thaliana
Sucrose Synthase 1 (EC:2.4.1.13), also known as the Sucrose-UDP glucolsyltransferase 1 or simply Susy, is a reversible enzyme allowing the synthesis or the degradation of Sucrose in Arabidopsis thaliana. It belongs to the Glycosyltransferase subfamily 4 (GT4).
| |||||||||||
References
• UniProt entry: P49040
• Brenda entry : 2.4.1.13
- ↑ Salerno GL, Curatti L Origin of sucrose metabolism in higher plants: when, how and why? Trends Plant Sci. 2003 Feb
- ↑ Baroja-Fernández, E., Muñoz, F.J., Saikusa, T., Rodríguez-López, M., Akazawa, T. and Pozueta-Romero, J. Sucrose synthase catalyzes the de novo production of ADPglucose linked to starch biosynthesis in heterotrophic tissues of plants. Plant Cell Physiol.
- ↑ Salerno GL, Curatti L. Origin of sucrose metabolism in higher plants: when, how and why? Trends Plant Sci. 2003 Feb
- ↑ Determination of structural requirements and probable regulatory effectors for membrane association of maize sucrose synthase 1. Hardin SC, Duncan KA, Huber SC. Plant Physiol. 2006
- ↑ Phosphorylation of sucrose synthase at serine 170: occurrence and possible role as a signal for proteolysis. Hardin SC, Tang GQ, Scholz A, Holtgraewe D, Winter H, Huber SC. Plant J. 2003