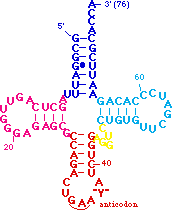We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Transfer RNA tour
From Proteopedia
(Difference between revisions)
m |
m |
||
| Line 1: | Line 1: | ||
| - | == | + | ==tRNA== |
<StructureSection load='1tra' size='400' side='right' caption='phe-tRNA [[1tra]]' scene='72/725890/Trna_overview/1'> | <StructureSection load='1tra' size='400' side='right' caption='phe-tRNA [[1tra]]' scene='72/725890/Trna_overview/1'> | ||
Source <ref>PMID:790568</ref> | Source <ref>PMID:790568</ref> | ||
| Line 22: | Line 22: | ||
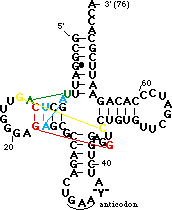
The red residues are a base triple in which <B><FONT COLOR="#ff0000">7-methyl-G46</FONT></B> from the <B>variable loop</B> H-bonds to the <B><FONT COLOR="#ff0000">G22-C13</FONT></B> base pair of the <B>D stem</B>. This helps dock the <B>variable loop</B> onto the <B>D-stem</B>. | The red residues are a base triple in which <B><FONT COLOR="#ff0000">7-methyl-G46</FONT></B> from the <B>variable loop</B> H-bonds to the <B><FONT COLOR="#ff0000">G22-C13</FONT></B> base pair of the <B>D stem</B>. This helps dock the <B>variable loop</B> onto the <B>D-stem</B>. | ||
| + | |||
| + | <B><FONT COLOR="#008aff">A9</FONT></B> is also involved in another type of tertiary interaction: it is <scene name='72/725890/Trna_tertwire/1'>intercalated between bases</scene> <B><FONT COLOR="#ff0000">7-methyl-G46</FONT></B> and <B>G45</B>. In order to make room between these bases for <B><FONT COLOR="#008aff">A9</FONT></B> the backbone is extended by a C2' endo sugar pucker at <B><FONT COLOR="#ff0000">7-methyl-G46</FONT></B>. | ||
| + | |||
===U-turns=== | ===U-turns=== | ||
Revision as of 20:29, 28 February 2016
tRNA
| |||||||||||
See Also
- Z-DNA model tour and Z-DNA
- B-DNA tour
- A-RNA tour
- A more general overview will be found at DNA.
- Forms of DNA shows a side-by-side comparison of A, B, and Z forms of DNA.
- An interactive tutorial on DNA Structure, disponible también en español and eight other languages.
References
JSmol in Proteopedia [2] or to the article describing Jmol [3] to the rescue.
- ↑ Quigley GJ, Rich A. Structural domains of transfer RNA molecules. Science. 1976 Nov 19;194(4267):796-806. PMID:790568
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644