This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1176
From Proteopedia
(Difference between revisions)
| Line 6: | Line 6: | ||
==Introduction== | ==Introduction== | ||
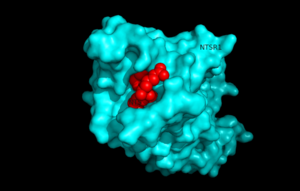
| - | [[Image:surfaceprotein.png |300 px|below|thumb|View of NTSR1 protein interacting with NTS ligand]] | + | [[Image:surfaceprotein.png |300 px|below|thumb|Figure 1.View of NTSR1 protein interacting with NTS ligand]] |
Neurotensin receptor 1 (NTSR1) is a G-protein coupled receptor (GPCR) that binds to the 13 amino acid peptide, neurotensin. Studies determining the structure of NTSR1 crystallized the GPCR bound with the C-terminus of its tridecapeptide ligand, <scene name='72/721548/Neurotensin/3'>NTS(8-13)</scene> because it has a higher potency and efficacy than its full-length counterpart. NTSR1 is a class A GPCR, and like all G-proteins, consists of an extracellular binding domain along with 7 transmembrane helices. Along with the ligand binding pocket at the top of the protein, NTSR1 also contains an allosteric Na+ ion binding pocket underneath. NTS binds to NTSR1, leading to a conformational change of the protein and modulation of second messengers. NTS has been shown to have a variety of biological activities including a role in the leptin signalling pathways, tumor growth, and dopamine regulation. The majority of effects of NTS are mediated through NTSR1. Research of the structure of NTSR1 has focused on the differences between its active and active-like states. | Neurotensin receptor 1 (NTSR1) is a G-protein coupled receptor (GPCR) that binds to the 13 amino acid peptide, neurotensin. Studies determining the structure of NTSR1 crystallized the GPCR bound with the C-terminus of its tridecapeptide ligand, <scene name='72/721548/Neurotensin/3'>NTS(8-13)</scene> because it has a higher potency and efficacy than its full-length counterpart. NTSR1 is a class A GPCR, and like all G-proteins, consists of an extracellular binding domain along with 7 transmembrane helices. Along with the ligand binding pocket at the top of the protein, NTSR1 also contains an allosteric Na+ ion binding pocket underneath. NTS binds to NTSR1, leading to a conformational change of the protein and modulation of second messengers. NTS has been shown to have a variety of biological activities including a role in the leptin signalling pathways, tumor growth, and dopamine regulation. The majority of effects of NTS are mediated through NTSR1. Research of the structure of NTSR1 has focused on the differences between its active and active-like states. | ||
| Line 33: | Line 33: | ||
===Schizophrenia Research=== | ===Schizophrenia Research=== | ||
| - | The | + | The '''[http://www.schizophreniaforum.org/for/curr/AbiDargham/ dopamine hypothesis]''' states that having hyperdopamine levels may lead to schizophrenic symptoms. It has been shown that NTSR1 caused a blockade which inhibited firing in dopaminergic cells. It is believed that NTSR1 could therefore be used as a therapeutic for treating schizophrenia.Although this research is promising, the secondary effects were too extreme and the trial was discontinued.This is a pathway where NTSR1 research is focused on that could lead to ground breaking advances in treating schizophrenia. |
==Structural Research== | ==Structural Research== | ||
Revision as of 17:31, 1 April 2016
| This Sandbox is Reserved from Jan 11 through August 12, 2016 for use in the course CH462 Central Metabolism taught by R. Jeremy Johnson at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1160 through Sandbox Reserved 1184. |
To get started:
More help: Help:Editing |
Rattus norevegicus NTSR1
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644