This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
TEM1 Class Antibiotic Resistance Proteins
From Proteopedia
(Difference between revisions)
| Line 28: | Line 28: | ||
== Function == | == Function == | ||
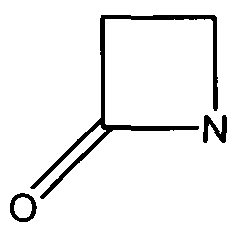
The β-Lactamases has two domains- an alpha helix and a beta sheet of five antiparallel strands, which surround the alpha helix. The catalytic region is known as the oxyanion pocket that occurs between the N-terminus of alpha helix H2 and B3 beta sheet edge. The first step is the acylation of Ser70, one of the catalytic sites. This forms a high-energy acyl-enzyme intermediate, which is then deacylated. This is when the acyl-enzyme is hydrolyzed. The β-Lactam compound has been split and released. The rate determining step is either the acylation or deacylation depending on the antibiotic. However, the opening of the β-Lactam active site increases its chance of inhibition. One of the ways to inhibit β-Lactamases is point mutations in the catalytic region that affect the specificity and catalysis of β-Lactamases enzymes, hindering their activity [4]. | The β-Lactamases has two domains- an alpha helix and a beta sheet of five antiparallel strands, which surround the alpha helix. The catalytic region is known as the oxyanion pocket that occurs between the N-terminus of alpha helix H2 and B3 beta sheet edge. The first step is the acylation of Ser70, one of the catalytic sites. This forms a high-energy acyl-enzyme intermediate, which is then deacylated. This is when the acyl-enzyme is hydrolyzed. The β-Lactam compound has been split and released. The rate determining step is either the acylation or deacylation depending on the antibiotic. However, the opening of the β-Lactam active site increases its chance of inhibition. One of the ways to inhibit β-Lactamases is point mutations in the catalytic region that affect the specificity and catalysis of β-Lactamases enzymes, hindering their activity [4]. | ||
| + | |||
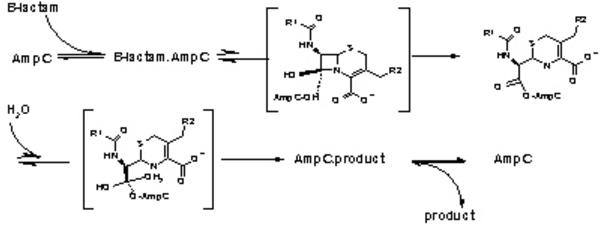
[[Image:Biochem_group_project-function.PNG]] | [[Image:Biochem_group_project-function.PNG]] | ||
| + | |||
Figure 3. The hydrolysis of β-Lactamases where AmpC is the enzyme. There are the three main steps shown with their respective high energy intermediates. (Image Credit: Shoichet Lab in UCSF) | Figure 3. The hydrolysis of β-Lactamases where AmpC is the enzyme. There are the three main steps shown with their respective high energy intermediates. (Image Credit: Shoichet Lab in UCSF) | ||
== Relevance == | == Relevance == | ||
| Line 46: | Line 48: | ||
4. Fonze, E.; Charlier, P.; To'th, Y.; Vermeire, M.; Raquet, X.; Dubus, A.; Frere, J. M. (1995) TEM1 beta-lactamase structure solved by molecular replacement and refined structure of the S235A mutant. ''Acta Crystallogr. D Biol. Crystallogr.'' ''51'', 682-694. | 4. Fonze, E.; Charlier, P.; To'th, Y.; Vermeire, M.; Raquet, X.; Dubus, A.; Frere, J. M. (1995) TEM1 beta-lactamase structure solved by molecular replacement and refined structure of the S235A mutant. ''Acta Crystallogr. D Biol. Crystallogr.'' ''51'', 682-694. | ||
| - | 5. Salverda, M. L.; Visser, J. A. G. D.; Barlow, M. Natural Evolution of TEM-1 β-Lactamase: Experimental Reconstruction and Clinical Relevance. FEMS Microbiology Reviews FEMS Microbiol Rev. | + | 5. Salverda, M. L.; Visser, J. A. G. D.; Barlow, M. (2010) Natural Evolution of TEM-1 β-Lactamase: Experimental Reconstruction and Clinical Relevance. ''FEMS Microbiology Reviews FEMS Microbiol Rev.'' ''34'', 1015–1036. |
| - | 6. Neuwirth, C.; Madec, S.; Siebor, E.; Pechinot, A.; Duez, J.-M.; Pruneaux, M.; Fouchereau-Peron, M.; Kazmierczak, A.; Labia, R. TEM-89 Beta -Lactamase Produced by a Proteus Mirabilis Clinical Isolate: New Complex Mutant (CMT 3) with Mutations in Both TEM-59 (IRT-17) and TEM-3. Antimicrobial Agents and Chemotherapy | + | 6. Neuwirth, C.; Madec, S.; Siebor, E.; Pechinot, A.; Duez, J.-M.; Pruneaux, M.; Fouchereau-Peron, M.; Kazmierczak, A.; Labia, R. (2010) TEM-89 Beta -Lactamase Produced by a Proteus Mirabilis Clinical Isolate: New Complex Mutant (CMT 3) with Mutations in Both TEM-59 (IRT-17) and TEM-3. ''Antimicrobial Agents and Chemotherapy'' ''45'', 3591–3594. |
| - | 7. Elander, R. P. Industrial Production of β-Lactam Antibiotics. Appl Microbiol Biotechnol Applied Microbiology and Biotechnology. | + | 7. Elander, R. P. (2003) Industrial Production of β-Lactam Antibiotics. ''Appl Microbiol Biotechnol Applied Microbiology and Biotechnology''. ''61'', 385–392. |
8. Lenfant, F.; Labia, R.; Masson, J. (1991) Replacement of lysine 234 affects transition state stabilization in the active site of ß-lactamase TEM1. ''J. Biol. Chem.'' ''266'', 17187-17194. | 8. Lenfant, F.; Labia, R.; Masson, J. (1991) Replacement of lysine 234 affects transition state stabilization in the active site of ß-lactamase TEM1. ''J. Biol. Chem.'' ''266'', 17187-17194. | ||
9. Doucet, N.; Savard, P. -.; Pelletier, J. N.; Gagné, S. M. (2007) NMR investigation of Tyr105 mutants in TEM-1 ß-lactamase: Dynamics are correlated with function. ''J. Biol. Chem.'' ''282'', 21448-21459. | 9. Doucet, N.; Savard, P. -.; Pelletier, J. N.; Gagné, S. M. (2007) NMR investigation of Tyr105 mutants in TEM-1 ß-lactamase: Dynamics are correlated with function. ''J. Biol. Chem.'' ''282'', 21448-21459. | ||
Revision as of 21:26, 27 April 2016
| |||||||||||
Proteopedia Page Contributors and Editors (what is this?)
Kenna Salvatore, Matt O'Malley, Ryan Hunter Wilson, Riley Culhane, Michal Harel