This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Urease accessory protein
From Proteopedia
(Difference between revisions)
| Line 6: | Line 6: | ||
== Structural highlights == | == Structural highlights == | ||
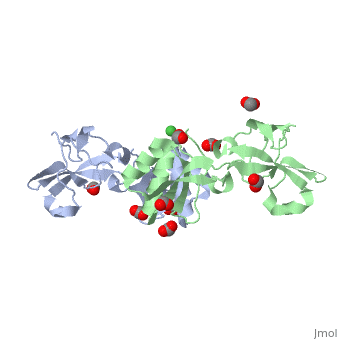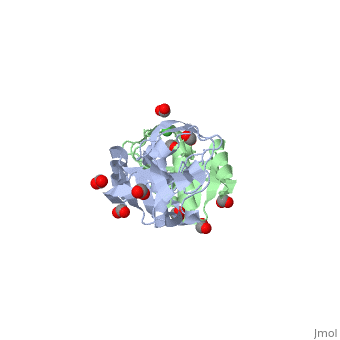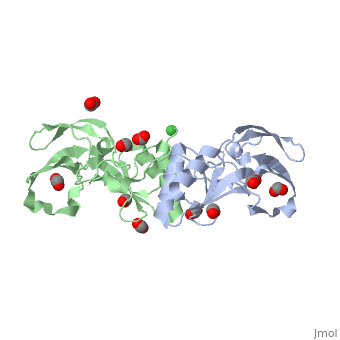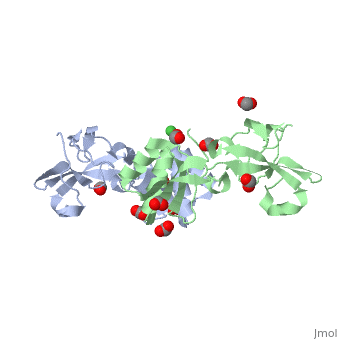
| + | The Ni+2 ion coordinates six ligands which include 3 His residues belonging to the two monomers, Glu from a symmetry-related monomer, water molecule and an unidentified ligand which may be a His from an unobserved part of the chain<ref>PMID:22010876</ref> | ||
</StructureSection> | </StructureSection> | ||
Revision as of 08:13, 8 December 2016
| |||||||||||
3D structures of urease accessory protein
Updated on 08-December-2016
References
- ↑ Witte CP, Isidore E, Tiller SA, Davies HV, Taylor MA. Functional characterisation of urease accessory protein G (ureG) from potato. Plant Mol Biol. 2001 Jan;45(2):169-79. PMID:11289508
- ↑ Banaszak K, Martin-Diaconescu V, Bellucci M, Zambelli B, Rypniewski W, Maroney MJ, Ciurli S. Crystallographic and X-ray absorption spectroscopic characterization of Helicobacter pylori UreE bound to Ni2+ and Zn2+ reveal a role for the disordered C-terminal arm in metal trafficking. Biochem J. 2011 Oct 20. PMID:22010876 doi:10.1042/BJ20111659