This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sucrase-isomaltase
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
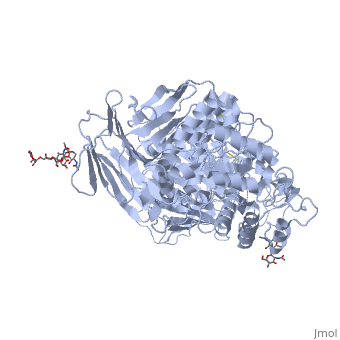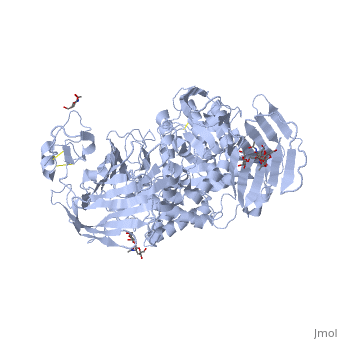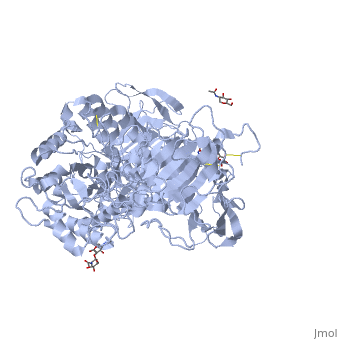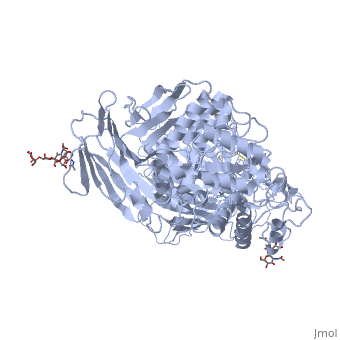
| - | < | + | <StructureSection load='3lpo' size='350' side='right' scene='' caption='Human glycosylated sucrase-isomaltase N-terminal domain (PDB code [[3lpo]])'> |
| - | + | ||
== Function == | == Function == | ||
| Line 45: | Line 44: | ||
-The gene on which SI processes occur is located on chromosome 3 and is composed of 48 exons. | -The gene on which SI processes occur is located on chromosome 3 and is composed of 48 exons. | ||
| + | </StructureSection> | ||
==3D structures of surase-isomaltase== | ==3D structures of surase-isomaltase== | ||
Revision as of 11:06, 4 June 2017
| |||||||||||
3D structures of surase-isomaltase
Updated on 04-June-2017
3lpo - hSI N-terminal - human
3lpp - hSI N-terminal + kotalanol
4m56 - BsSI + glucose - Bacillus subtilis
4m8u, 4maz, 4mb1 - BsSI (mutant)
References
- ↑ http://www.siumed.edu/~dking2/erg/gicells.htm
- ↑ http://www.uniprot.org/uniprot/P14410
- ↑ Maureen Barlow Pugh, ed. (2000). Stedman's Medical Dictionary (27th ed.). Baltimore, Maryland, USA: Lippincott Williams & Wilkins. p. 65. ISBN 978-0-683-40007-6.
- ↑ "SI sucrase-isomaltase (alpha-glucosidase) [Homo sapiens (human)] - Gene - NCBI"
- ↑ Naim HY, Sterchi EE, Lentze MJ (1988). "Biosynthesis of the human sucrase-isomaltase complex. Differential O-glycosylation of the sucrase subunit correlates with its position within the enzyme complex". J. Biol. Chem. 263 (15): 7242–53. PMID 3366777
- ↑ http://www.jbc.org/content/254/6/1821.full.pdf
- ↑ http://onlinelibrary.wiley.com/doi/10.1111/j.1432-1033.1977.tb11335.x/epdf
- ↑ http://www.jbc.org/content/251/11/3250.full.pdf
- ↑ http://www.iffgd.org/site/gi-disorders/other/csid
- ↑ Sim L, Willemsma C, Mohan S, Naim HY, Pinto BM, Rose DR (2010). "Structural basis for substrate selectivity in human maltase-glucoamylase and sucrase-isomaltase N-terminal domains". J. Biol. Chem. 285 (23): 17763–70. doi:10.1074/jbc.M109.078980. PMC 2878540. PMID 20356844