This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Luke Edward Severinac/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 10: | Line 10: | ||
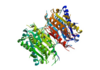
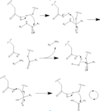
Caspase 6 is a part of the cystine aspartic protease family that cleaves proteins at the TEVD sequence. in its monomeric form with protein ligand bound, its catalytic residues are <scene name='75/752344/His121/1'>Histidine 121</scene>, <scene name='75/752344/Glu123/1'>Glutamate 123</scene>, and <scene name='75/752344/Cys163/1'>Cystine 163</scene>.[[Image:Mechanism caspase 6.PNG|100 px|left|thumb|Cystine Aspartase mechanism]] Together, these residues form a <scene name='75/752344/Caspase-6_catalytic_triad_real/1'>catalytic triad</scene> that cleaves a <scene name='75/752344/Protein_ligand/1'>protein ligand</scene>. | Caspase 6 is a part of the cystine aspartic protease family that cleaves proteins at the TEVD sequence. in its monomeric form with protein ligand bound, its catalytic residues are <scene name='75/752344/His121/1'>Histidine 121</scene>, <scene name='75/752344/Glu123/1'>Glutamate 123</scene>, and <scene name='75/752344/Cys163/1'>Cystine 163</scene>.[[Image:Mechanism caspase 6.PNG|100 px|left|thumb|Cystine Aspartase mechanism]] Together, these residues form a <scene name='75/752344/Caspase-6_catalytic_triad_real/1'>catalytic triad</scene> that cleaves a <scene name='75/752344/Protein_ligand/1'>protein ligand</scene>. | ||
===Zinc Exoside=== | ===Zinc Exoside=== | ||
| + | |||
| + | =='''Activation of Caspase-6'''== | ||
| + | |||
| + | ===Subunits involved in activation=== | ||
| + | |||
| + | ===Self cleavage=== | ||
==Inhbition== | ==Inhbition== | ||
Revision as of 01:09, 2 April 2017
Caspase 6 in Homo Sapiens
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2966951/ (self cleavage article)
http://www.rcsb.org/pdb/explore/explore.do?structureId=2WDP (this is the non-self cleaved protien)