This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Luke Edward Severinac/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
Found at high concentrations in the brain and bordering tissues, Caspase-6 has been implicated in several neurological diseases including Alzheimer's and dementia[http://www.alz.org/]<ref name="ActiveRegofCasp6andNDdisease">PMID: 25340928 </ref>. It's primarily involved in apoptosis through a largely ambiguous mechanism. It is classified as an endopeptidase[https://en.wikipedia.org/wiki/Endopeptidase] as it cleaves an internal peptide bond of its substrate. It has relatively low specificity in the binding site which allows for a variety of substrates, including other caspase enzymes and neuronal proteins to bind<ref name="ZincMediatedCasp6">PMID: 22891250 </ref>. Furthermore, it is a part of the cysteine-aspartate family[https://en.wikipedia.org/wiki/Caspase], which have these critical amino acid residues in the active site of the enzyme. Caspase-6 has both an inactive zinc-bound conformation and an active ligand-bound conformation, which are largely regulated by variations in zinc concentration<ref name="ZincMediatedCasp6">PMID: 22891250 </ref>. | Found at high concentrations in the brain and bordering tissues, Caspase-6 has been implicated in several neurological diseases including Alzheimer's and dementia[http://www.alz.org/]<ref name="ActiveRegofCasp6andNDdisease">PMID: 25340928 </ref>. It's primarily involved in apoptosis through a largely ambiguous mechanism. It is classified as an endopeptidase[https://en.wikipedia.org/wiki/Endopeptidase] as it cleaves an internal peptide bond of its substrate. It has relatively low specificity in the binding site which allows for a variety of substrates, including other caspase enzymes and neuronal proteins to bind<ref name="ZincMediatedCasp6">PMID: 22891250 </ref>. Furthermore, it is a part of the cysteine-aspartate family[https://en.wikipedia.org/wiki/Caspase], which have these critical amino acid residues in the active site of the enzyme. Caspase-6 has both an inactive zinc-bound conformation and an active ligand-bound conformation, which are largely regulated by variations in zinc concentration<ref name="ZincMediatedCasp6">PMID: 22891250 </ref>. | ||
| - | Caspase-6 is an endopeptidase [https://en.wikipedia.org/wiki/Endopeptidase] involved in apoptosis. In terms of its catalytic function, it is a part of the cysteine-aspartate family [https://en.wikipedia.org/wiki/Caspase]. Before Caspase-6 becomes functional and active, the enzyme exists as a procaspase, also known as a zymogen [https://en.wikipedia.org/wiki/Zymogen][6]. In solution, two zymogens are associated together, forming a homodimer. Zymogen activation, the process by which Caspase-6 becomes active, is largely conserved across the caspase family. | + | Caspase-6 is an endopeptidase [https://en.wikipedia.org/wiki/Endopeptidase] involved in apoptosis. In terms of its catalytic function, it is a part of the cysteine-aspartate family [https://en.wikipedia.org/wiki/Caspase]. Before Caspase-6 becomes functional and active, the enzyme exists as a procaspase, also known as a zymogen [https://en.wikipedia.org/wiki/Zymogen][6]. In solution, two zymogens are associated together, forming a homodimer. Zymogen activation, the process by which Caspase-6 becomes active, is largely conserved across the caspase family. |
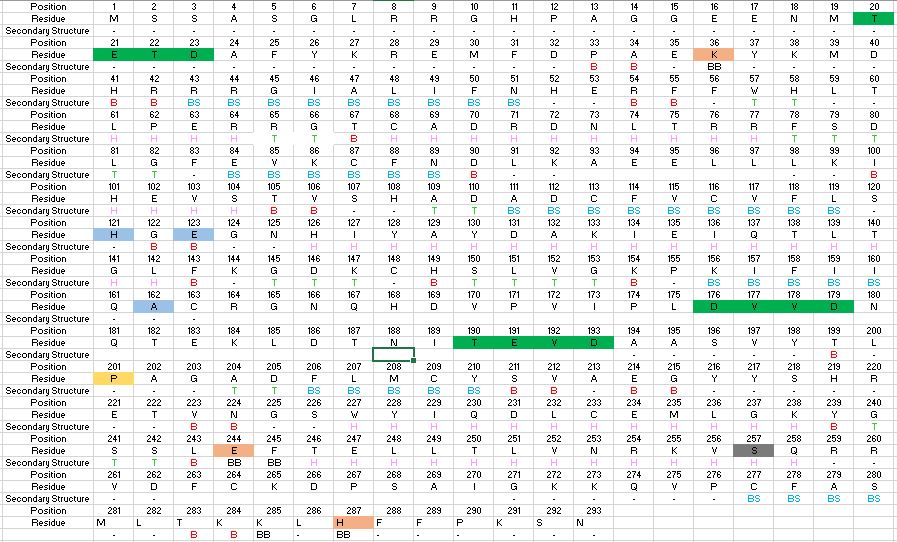
| - | + | However, Caspase-6 is unique in that it becomes active through self-cleavage rather than cleavage by a separate enzyme. Each zymogen of the unprocessed enzyme contains a small subunit consisting of two helices and large subunit consisting of three helices, a prodomain, as well as an intersubunit linker. The helices surround a beta sheet core. In order to become active, the intersubunit linker is bound to the active site of Caspase-6, where it is then cleaved. After cleavage, the four processed subunits, two originating from each zymogen, remain closely associated together through intermolecular forces, forming a dimer of dimers. | |
[[Image:Caspase-6 protein.jpg|100 px|left|thumb|Caspase-6 Protein]] | [[Image:Caspase-6 protein.jpg|100 px|left|thumb|Caspase-6 Protein]] | ||
Revision as of 23:33, 17 April 2017
Caspase-6 in Homo sapiens
| |||||||||||
References
- ↑ 1.0 1.1 1.2 Wang XJ, Cao Q, Zhang Y, Su XD. Activation and regulation of caspase-6 and its role in neurodegenerative diseases. Annu Rev Pharmacol Toxicol. 2015;55:553-72. doi:, 10.1146/annurev-pharmtox-010814-124414. Epub 2014 Oct 17. PMID:25340928 doi:http://dx.doi.org/10.1146/annurev-pharmtox-010814-124414
- ↑ 2.0 2.1 Velazquez-Delgado EM, Hardy JA. Zinc-Mediated Allosteric Inhibition of Caspase-6. J Biol Chem. 2012 Aug 13. PMID:22891250 doi:http://dx.doi.org/10.1074/jbc.M112.397752
- ↑ Wang XJ, Cao Q, Liu X, Wang KT, Mi W, Zhang Y, Li LF, Leblanc AC, Su XD. Crystal structures of human caspase 6 reveal a new mechanism for intramolecular cleavage self-activation. EMBO Rep. 2010 Oct 1. PMID:20890311 doi:10.1038/embor.2010.141
- ↑ Velazquez-Delgado EM, Hardy JA. Phosphorylation regulates assembly of the caspase-6 substrate-binding groove. Structure. 2012 Apr 4;20(4):742-51. Epub 2012 Apr 3. PMID:22483120 doi:10.1016/j.str.2012.02.003