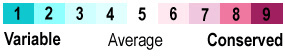This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1057
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
==Introduction== | ==Introduction== | ||
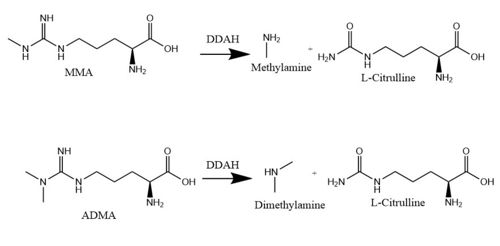
| - | Dimethylarginine Dimethyaminohydrolase <span class="plainlinks">[http://www.chem.qmul.ac.uk/iubmb/enzyme/EC3/5/3/18.html EC 3.5.3.18]</span> (commonly known as DDAH) is a member of the <span class="plainlinks">[https://en.wikipedia.org/wiki/Hydrolase hydrolase]</span> family of enzymes which use water to break down molecules <ref name="palm">Palm F, Onozato ML, Luo Z, Wilcox CS. Dimethylarginine dimethylaminohydrolase (DDAH): expression, regulation, and function in the cardiovascular and renal systems. American Journal of Physiology. 2007 Dec 1;293(6):3227-3245. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pubmed/17933965 17933965]</span> doi:<span class="plainlinks">[http://ajpheart.physiology.org/content/293/6/H3227 10.1152/ajpheart.00998.2007]</span></ref>. Additionally, DDAH is a <span class="plainlinks">[https://en.wikipedia.org/wiki/Nitric_oxide_synthase nitric oxide synthase (NOS)]</span> regulator. It metabolizes free arginine derivatives, namely <span class="plainlinks">[https://en.wikipedia.org/wiki/Asymmetric_dimethylarginine N<sup>Ѡ</sup>,N<sup>Ѡ</sup>-dimethyl-L-arginine (ADMA)]</span> and <span class="plainlinks">[https://en.wikipedia.org/wiki/Methylarginine N<sup>Ѡ</sup>-methyl-L-arginine (MMA)]</span> which competitively inhibit NOS <ref name="tran">Tran CTL, Leiper JM, Vallance P. The DDAH/ADMA/NOS pathway. Atherosclerosis Supplements. 2003 Dec;4(4):33-40. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pubmed/14664901 14664901]</span> doi:<span class="plainlinks">[http://www.sciencedirect.com/science/article/pii/S1567568803000321 10.1016/S1567-5688(03)00032-1]</span></ref>. DDAH converts MMA or ADMA to two products: <span class="plainlinks">[https://en.wikipedia.org/wiki/Citrulline L-citrulline]</span> and an amine <ref name="frey">Frey D, Braun O, Briand C, Vasak M, Grutter MG. Structure of the mammalian NOS regulator dimethylarginine dimethylaminohydrolase: a basis for the design of specific inhibitors. Structure. 2006 May;14(5):901-911. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pubmed/ | + | Dimethylarginine Dimethyaminohydrolase <span class="plainlinks">[http://www.chem.qmul.ac.uk/iubmb/enzyme/EC3/5/3/18.html EC 3.5.3.18]</span> (commonly known as DDAH) is a member of the <span class="plainlinks">[https://en.wikipedia.org/wiki/Hydrolase hydrolase]</span> family of enzymes which use water to break down molecules <ref name="palm">Palm F, Onozato ML, Luo Z, Wilcox CS. Dimethylarginine dimethylaminohydrolase (DDAH): expression, regulation, and function in the cardiovascular and renal systems. American Journal of Physiology. 2007 Dec 1;293(6):3227-3245. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pubmed/17933965 17933965]</span> doi:<span class="plainlinks">[http://ajpheart.physiology.org/content/293/6/H3227 10.1152/ajpheart.00998.2007]</span></ref>. Additionally, DDAH is a <span class="plainlinks">[https://en.wikipedia.org/wiki/Nitric_oxide_synthase nitric oxide synthase (NOS)]</span> regulator. It metabolizes free arginine derivatives, namely <span class="plainlinks">[https://en.wikipedia.org/wiki/Asymmetric_dimethylarginine N<sup>Ѡ</sup>,N<sup>Ѡ</sup>-dimethyl-L-arginine (ADMA)]</span> and <span class="plainlinks">[https://en.wikipedia.org/wiki/Methylarginine N<sup>Ѡ</sup>-methyl-L-arginine (MMA)]</span> which competitively inhibit NOS <ref name="tran">Tran CTL, Leiper JM, Vallance P. The DDAH/ADMA/NOS pathway. Atherosclerosis Supplements. 2003 Dec;4(4):33-40. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pubmed/14664901 14664901]</span> doi:<span class="plainlinks">[http://www.sciencedirect.com/science/article/pii/S1567568803000321 10.1016/S1567-5688(03)00032-1]</span></ref>. DDAH converts MMA or ADMA to two products: <span class="plainlinks">[https://en.wikipedia.org/wiki/Citrulline L-citrulline]</span> and an amine <ref name="frey">Frey D, Braun O, Briand C, Vasak M, Grutter MG. Structure of the mammalian NOS regulator dimethylarginine dimethylaminohydrolase: a basis for the design of specific inhibitors. Structure. 2006 May;14(5):901-911. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pubmed/16698551]</span> doi:<span class="plainlinks">[http://www.sciencedirect.com/science/article/pii/S0969212606001717 10.1016/j.str.2006.03.006]</span></ref> (Figure 1). DDAH is expressed in the cytosol of cells in humans, mice, rates, sheep, cattle, and bacteria <ref name="palm" />. DDAH activity has been localized mainly to the brain, kidney, pancreas, and liver in these organisms. Presented in this page is information from DDAH isoform 1 (DDAH-1); however, there are two different isoforms <ref name="frey" />. |
[[Image:DDAH mechanism.jpg|500 px|center|thumb|'''Figure 1.''' The normal DDAH mechanism]] | [[Image:DDAH mechanism.jpg|500 px|center|thumb|'''Figure 1.''' The normal DDAH mechanism]] | ||
| Line 10: | Line 10: | ||
==General Structure== | ==General Structure== | ||
| - | DDAH’s <scene name='69/694225/Secondary_structure_colored/3'>secondary structure</scene> has a <scene name='69/694225/Prop_domains/2'>propeller-like fold</scene> which is characteristic of the superfamily of <span class="plainlinks">[https://en.wikipedia.org/wiki/Arginine:glycine_amidinotransferase L-arginine/glycine amidinotransferases]</span> <ref name="humm">Humm A, Fritsche E, Mann K, Göhl M, Huber R. Recombinant expression and isolation of human L-arginine:glycine amidinotransferase and identification of its active-site cysteine residue. Biochemical Journal. 1997 March 15;322(3):771-776. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1218254/ 9148748]</span> doi:<span class="plainlinks">[http://www.biochemj.org/content/322/3/771 10.1042/bj3220771]</span></ref>. This five-stranded | + | DDAH’s <scene name='69/694225/Secondary_structure_colored/3'>secondary structure</scene> has a <scene name='69/694225/Prop_domains/2'>propeller-like fold</scene> which is characteristic of the superfamily of <span class="plainlinks">[https://en.wikipedia.org/wiki/Arginine:glycine_amidinotransferase L-arginine/glycine amidinotransferases]</span> <ref name="humm">Humm A, Fritsche E, Mann K, Göhl M, Huber R. Recombinant expression and isolation of human L-arginine:glycine amidinotransferase and identification of its active-site cysteine residue. Biochemical Journal. 1997 March 15;322(3):771-776. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1218254/ 9148748]</span> doi:<span class="plainlinks">[http://www.biochemj.org/content/322/3/771 10.1042/bj3220771]</span></ref>. This five-stranded <span class="plainlinks">[https://en.wikipedia.org/wiki/Beta-propeller propeller]</span> contains five repeats of a ββαβ motif <ref name="frey" />. These motifs in DDAH form a <scene name='75/752351/Ddah_water_pore/9'>channel</scene> filled with water molecules (red spheres). Lys174 and Glu77 form a <scene name='75/752351/Ddah_salt_bridge/4'>salt bridge</scene> in the channel that forms the bottom of the <scene name='75/752351/Ddah_active_site/1'>active site</scene> for the protein. One side of the channel is a <scene name='75/752351/Ddah_water_pore/3'>water-filled pore</scene>, whereas the other side is the active site cleft <ref name="frey" />. |
===Lid Region=== | ===Lid Region=== | ||
| - | Amino acids 25-36 of DDAH constitute the loop region of the protein which is more commonly known as the lid region <ref name="frey" /> | + | Amino acids 25-36 of DDAH constitute the flexible |
| + | <scene name='75/752351/Lid_focus/1'>loop region</scene> of the protein which is more commonly known as the lid region <ref name="frey" />. The lid is what allows the active site to be exposed to substrate binding or not. Studies have shown crystal structures of the lid at <scene name='69/694225/Open_surface/5'>open</scene>and <scene name='69/694225/Closed_surface/4'>closed</scene> conformations. In the open conformation, the lid forms an alpha helix and the amino acid <scene name='69/694225/Lid_helix/1'>Leu29</scene> is moved so it does not interact with the active site. This allows the active site to be vulnerable to attack. When the lid is closed, a specific <scene name=’75/752351/Hbond_leu29/1’>hydrogen bond</scene> can form between the Leu29 carbonyl and the amino group on bound molecule. This stabilizes this complex. The Leu29 is then <scene name='69/694225/Closed_lid_zn9/3'>blocking</scene> the active site entrance <ref name="frey" />. Opening and closing the lid takes place faster than the actual reaction in the active site <ref name="rasheed">Rasheed M, Richter C, Chisty LT, Kirkpatrick J, Blackledge M, Webb MR, Driscoll PC. Ligand-dependent dynamics of the active site lid in bacterial Dimethyarginine Dimethylaminohydrolase. Biochemistry. 2014 Feb 18;53:1092-1104. PMCID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3945819/ PMC3945819]</span> doi:<span class="plainlinks">[http://pubs.acs.org/doi/abs/10.1021/bi4015924 10.1021/bi4015924]</span></ref>. This suggests that the <span class="plainlinks">[https://en.wikipedia.org/wiki/Rate-determining_step rate-limiting step]</span> of this reaction is not the lid movement but is the actual chemistry happening to the substrate in the active site of DDAH <ref name="rasheed" />. | ||
====Lid Region Conservation==== | ====Lid Region Conservation==== | ||
| Line 27: | Line 28: | ||
====Active Site Conservation==== | ====Active Site Conservation==== | ||
| - | Active sites of DDAH from different organisms are similar. Amino acids involved in the chemical mechanism of creating products are also <scene name='69/694225/Evolutionary_conservation/ | + | Active sites of DDAH from different organisms are similar. Amino acids involved in the chemical mechanism of creating products are also <scene name='69/694225/Evolutionary_conservation/2'>conserved</scene> (Figure 4). |
[[Image:ColorKey ConSurf NoYellow NoGray.gif|400px|right|thumb|'''Figure 4.''' Color key for DDAH conservation]] | [[Image:ColorKey ConSurf NoYellow NoGray.gif|400px|right|thumb|'''Figure 4.''' Color key for DDAH conservation]] | ||
| Line 35: | Line 36: | ||
=====Important residues in Zinc Binding===== | =====Important residues in Zinc Binding===== | ||
| - | It was found that Cys273, His172, Glu77, Asp78, and Asp 268 all play a role in the binding of Zn(II). <scene name='69/694225/ | + | It was found that Cys273, His172, Glu77, Asp78, and Asp 268 all <scene name='69/694225/Active_site6hbonds/2'>play a role</scene> in the binding of Zn(II). <scene name='69/694225/Cys273_zn/1'>Cys273</scene> directly coordinates with the Zn(II) ion in the active site while the other significant residues stabilize the ion via hydrogen bonding interactions with water molecules in the active site. Depending on pH, His172 can <scene name='69/694225/Active_site_9/1'>change conformation</scene> and use the <span class="plainlinks">[https://en.wikipedia.org/wiki/Imidazole imidazole]</span> group to directly coordinate the Zn(II) ion. Cys273, which is conserved between bovine and humans, is the key active site residue that coordinates Zn(II) <ref name="frey" />. Zinc-cysteine complexes have been found to be important mediators of protein <span class="plainlinks">[https://en.wikipedia.org/wiki/Catalysis catalysis]</span>, regulation, and structure <ref name="pace">Pace NJ, Weerpana E. Zinc-binding cysteines: diverse functions and structural motifs. Biomolecules. 2014 June;4(2):419-434. PMCID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4101490/ 4101490]</span> doi:<span class="plainlinks">[http://www.mdpi.com/2218-273X/4/2/419/htm 10.3390/biom4020419]</span> </ref>. Cys273 and the water molecules stabilize the Zn(II) ion in a tetrahedral environment. The Zn(II) dissociation constant is 4.2 nM which is consistent with the nanomolar concentrations of Zn(II) in the cells, which provides more evidence for the regulatory use of Zn(II) by DDAH <ref name="pace" />. |
====Inhibitors==== | ====Inhibitors==== | ||
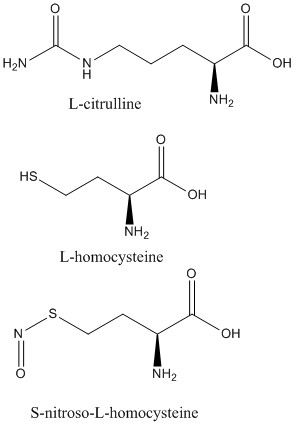
| - | <scene name='75/752351/Ddah_l-homocysteine/2'>L-homocysteine</scene> and <scene name='75/752351/Ddah_with_L-citrulline/4'>L-citrulline</scene> bind in the active site in the same orientation as MMA and ADMA to create the same <span class="plainlinks">[https://en.wikipedia.org/wiki/Intermolecular_force intermolecular bonds]</span> between them and DDAH <ref name="frey" /> (Figure 6). L-citrulline is a product of DDAH hydrolyzing ADMA and MMA, suggesting DDAH activity creates a <span class="plainlinks">[https://en.wikipedia.org/wiki/Negative_feedback negative feedback]</span> loop on itself. Both molecules enter the active site and cause DDAH to be in its closed lid formation. The α carbon on either molecule creates three salt bridges with DDAH: two with the guanidine group of Arg144 and one with the guanidine group Arg97. Another salt bridge is formed between the ligand and Asp72. The <scene name= | + | <scene name='75/752351/Ddah_l-homocysteine/2'>L-homocysteine</scene> and <scene name='75/752351/Ddah_with_L-citrulline/4'>L-citrulline</scene> bind in the active site in the same orientation as MMA and ADMA to create the same <span class="plainlinks">[https://en.wikipedia.org/wiki/Intermolecular_force intermolecular bonds]</span> between them and DDAH <ref name="frey" /> (Figure 6). L-citrulline is a product of DDAH hydrolyzing ADMA and MMA, suggesting DDAH activity creates a <span class="plainlinks">[https://en.wikipedia.org/wiki/Negative_feedback negative feedback]</span> loop on itself. Both molecules enter the active site and cause DDAH to be in its closed lid formation. The α carbon on either molecule creates three salt bridges with DDAH: two with the guanidine group of Arg144 and one with the guanidine group Arg97. Another salt bridge is formed between the ligand and Asp72. The molecules are stabilized in the active site by <scene name='75/752351/Hbond_leu29/2'>four hydrogen bonds</scene>: α carbon-amino group of the ligand to main chain carbonyls of Val267 and Leu29. Hydrogen bonds also form between the side chains of Asp78 and Glu77 with the ureido group of L-citrulline. |
Like L-homocysteine and L-citrulline, <scene name='75/752351/Ddah_s-nitroso-l-homocysteine/3'>S-nitroso-L-homocysteine</scene> binds and the lid region of DDAH is closed (Figure 6). When DDAH reacts with S-nitroso-L-homocysteine, a covalent product, N-thiosulfximide exist in the active site because of its binding to Cys273. N-thiosulfximide is stabilized by several salt bridges and hydrogen bonds. Arg144 and Arg97 stabilize the α carbon-carbonyl group via salt bridges, and Leu29, Val267, and Asp72 stabilize the α carbon-amino group by forming hydrogen bonds <ref name="frey" />. | Like L-homocysteine and L-citrulline, <scene name='75/752351/Ddah_s-nitroso-l-homocysteine/3'>S-nitroso-L-homocysteine</scene> binds and the lid region of DDAH is closed (Figure 6). When DDAH reacts with S-nitroso-L-homocysteine, a covalent product, N-thiosulfximide exist in the active site because of its binding to Cys273. N-thiosulfximide is stabilized by several salt bridges and hydrogen bonds. Arg144 and Arg97 stabilize the α carbon-carbonyl group via salt bridges, and Leu29, Val267, and Asp72 stabilize the α carbon-amino group by forming hydrogen bonds <ref name="frey" />. | ||
[[Image:L-citrulline, L-homocysteine, and S-nitroso-L-homocysteine.jpg|500px|center|thumb|'''Figure 6.''' Structures of DDAH inhibitors.]] | [[Image:L-citrulline, L-homocysteine, and S-nitroso-L-homocysteine.jpg|500px|center|thumb|'''Figure 6.''' Structures of DDAH inhibitors.]] | ||
Revision as of 00:58, 21 April 2017
Dimethylarginine Dimethylaminohydrolase
| |||||||||||
References
- ↑ 1.0 1.1 Palm F, Onozato ML, Luo Z, Wilcox CS. Dimethylarginine dimethylaminohydrolase (DDAH): expression, regulation, and function in the cardiovascular and renal systems. American Journal of Physiology. 2007 Dec 1;293(6):3227-3245. PMID:17933965 doi:10.1152/ajpheart.00998.2007
- ↑ 2.0 2.1 2.2 Tran CTL, Leiper JM, Vallance P. The DDAH/ADMA/NOS pathway. Atherosclerosis Supplements. 2003 Dec;4(4):33-40. PMID:14664901 doi:10.1016/S1567-5688(03)00032-1
- ↑ 3.00 3.01 3.02 3.03 3.04 3.05 3.06 3.07 3.08 3.09 3.10 3.11 3.12 3.13 3.14 3.15 3.16 3.17 3.18 3.19 3.20 3.21 3.22 3.23 Frey D, Braun O, Briand C, Vasak M, Grutter MG. Structure of the mammalian NOS regulator dimethylarginine dimethylaminohydrolase: a basis for the design of specific inhibitors. Structure. 2006 May;14(5):901-911. PMID:[1] doi:10.1016/j.str.2006.03.006
- ↑ Humm A, Fritsche E, Mann K, Göhl M, Huber R. Recombinant expression and isolation of human L-arginine:glycine amidinotransferase and identification of its active-site cysteine residue. Biochemical Journal. 1997 March 15;322(3):771-776. PMID:9148748 doi:10.1042/bj3220771
- ↑ 5.0 5.1 5.2 Rasheed M, Richter C, Chisty LT, Kirkpatrick J, Blackledge M, Webb MR, Driscoll PC. Ligand-dependent dynamics of the active site lid in bacterial Dimethyarginine Dimethylaminohydrolase. Biochemistry. 2014 Feb 18;53:1092-1104. PMCID:PMC3945819 doi:10.1021/bi4015924
- ↑ 6.0 6.1 Stone EM, Costello AL, Tierney DL, Fast W. Substrate-assisted cysteine deprotonation in the mechanism of Dimethylargininase (DDAH) from Pseudomonas aeruginosa. Biochemistry. 2006 May 2;45(17):5618-5630. PMID:16634643 doi:10.1021/bi052595m
- ↑ 7.0 7.1 Pace NJ, Weerpana E. Zinc-binding cysteines: diverse functions and structural motifs. Biomolecules. 2014 June;4(2):419-434. PMCID:4101490 doi:10.3390/biom4020419
- ↑ Janssen W, Pullamsetti SS, Cooke J, Weissmann N, Guenther A, Schermuly RT. The role of dimethylarginine dimethylaminohydrolase (DDAH) in pulmonary fibrosis. The Journal of Pathology. 2012 Dec 12;229(2):242-249. Epub 2013 Jan. PMID:23097221 doi:10.1002/path.4127
- ↑ Colasanti M, Suzuki H. The dual personality of NO. ScienceDirect. 2000 Jul 1;21(7):249-252. PMID:10979862 doi:10.1016/S0165-6147(00)01499-1
- ↑ Rassaf T, Feelisch M, Kelm M. Circulating NO pool: assessment of nitrite and nitroso species in blood and tissues. Free Rad. Biol. Med. 2004 Feb 15;36(4):413-422. PMID:14975444 doi:10.1016/j.freeradbiomed.2003.11.011
- ↑ Tsao PS, Cooke JP. Endothelial alterations in hypercholesterolemia: more than simply vasodilator dysfunction. Journal of Cardiovascular Pharmacology. 1998;32(3):48-53. PMID:9883748
- ↑ Vallance P, Leiper J. Blocking NO synthesis: how, where and why? Nat. Rev. Drug Discov. 2002 Dec;1(12):939-950. PMID:12461516 doi:10.1038/nrd960
Student Contributors
- Natalie Van Ochten
- Kaitlyn Enderle
- Colton Junod
3D Structures of Dimethylarginine Dimethylaminohydrolase
2C6Z L-citrulline bound to isoform 1
2CI1 S-nitroso-L-homocysteine bound to isoform 1
2CI3 crystal form 1
2CI4 crystal form 2
2CI5 L-homocysteine bound to isoform 1
2CI6 Zn (II) bound at low pH to isoform 1
2CI7 Zn (II) bound at high pH to isoform 1