This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1072
From Proteopedia
(Difference between revisions)
| Line 35: | Line 35: | ||
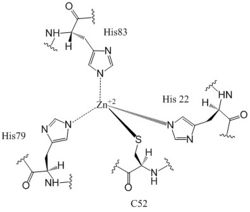
Most cells possess efficient Zinc uptake systems, as Zinc is a reactive Lewis Acid. Zinc binds incredibly tightly to this enzyme at subfemtomolar concentrations, attributing to why the enzyme has not been crystallized without Zinc present. The Zinc co-purified with the protein. Zinc allosterically inhibits the activity of enzyme DgcZ through two allosteric binding sites located on the CZB domain <sup>[1]</sup>. The inhibition prevents regulation of GGDEF domain function, the location of the active site. The CZB domain is folded into four anti-parallel α-helices as a 2-fold symmetric homodimer, with the N-terminus on the helix 𝝰4. The allosteric binding site includes a <scene name='69/694239/Zinc_binding_domain/4'>3His/1Cys</scene> motif that uses amino acids H22 of 𝝰1, C52 of 𝝰2, and H79 and H83 of 𝝰3, spanning three of the four alpha helices of the CZB domain and coordinating the Zinc residue in a tetrahedral fashion. For clarification, the entirety of 𝝰helix 2 on one monomer of CZB is not successfully crystallized after the Cys52 residue and is not the N-terminal residue. | Most cells possess efficient Zinc uptake systems, as Zinc is a reactive Lewis Acid. Zinc binds incredibly tightly to this enzyme at subfemtomolar concentrations, attributing to why the enzyme has not been crystallized without Zinc present. The Zinc co-purified with the protein. Zinc allosterically inhibits the activity of enzyme DgcZ through two allosteric binding sites located on the CZB domain <sup>[1]</sup>. The inhibition prevents regulation of GGDEF domain function, the location of the active site. The CZB domain is folded into four anti-parallel α-helices as a 2-fold symmetric homodimer, with the N-terminus on the helix 𝝰4. The allosteric binding site includes a <scene name='69/694239/Zinc_binding_domain/4'>3His/1Cys</scene> motif that uses amino acids H22 of 𝝰1, C52 of 𝝰2, and H79 and H83 of 𝝰3, spanning three of the four alpha helices of the CZB domain and coordinating the Zinc residue in a tetrahedral fashion. For clarification, the entirety of 𝝰helix 2 on one monomer of CZB is not successfully crystallized after the Cys52 residue and is not the N-terminal residue. | ||
| - | Zahringer et al. mutated Cys52 to Ala through <span class="plainlinks">[https://en.wikipedia.org/wiki/Site-directed_mutagenesis site-directed mutagenesis]</span>, resulting in a lack of coordination on α2. The cysteine residue is not essential for Zinc binding, as Zinc still coordinates to the three His residues with the Cys52Ala mutation, but α2 is free to move and expose the Zinc binding pocket. This exposure was found to lower the protein's affinity for zinc, as the mutation of cysteine to alanine increased the activity of the DgcZ. Using EDTA, Zinc can be removed from the CZB domain. When not coordinated to zinc, the CZB domain presumably adopts a conformation that straightens the <scene name='69/694239/Czbd_with_helices_labeled/ | + | Zahringer et al. mutated Cys52 to Ala through <span class="plainlinks">[https://en.wikipedia.org/wiki/Site-directed_mutagenesis site-directed mutagenesis]</span>, resulting in a lack of coordination on α2. The cysteine residue is not essential for Zinc binding, as Zinc still coordinates to the three His residues with the Cys52Ala mutation, but α2 is free to move and expose the Zinc binding pocket. This exposure was found to lower the protein's affinity for zinc, as the mutation of cysteine to alanine increased the activity of the DgcZ. Using EDTA, Zinc can be removed from the CZB domain. When not coordinated to zinc, the CZB domain presumably adopts a conformation that straightens the <scene name='69/694239/Czbd_with_helices_labeled/4'>α2 helix</scene>, shifting <scene name='69/694239/Hydrophobicity_int_residues/3'>hydrophobic residues</scene> on the α-helices into the center and the GGEEF domain into its productive conformation, increasing activity of DgcZ. Activity increases without Zinc due to activation of poly-GlcNAc production and biofilm formation, and maximal cyclic di-GMP production. |
=== Other Ligands === | === Other Ligands === | ||
Revision as of 01:42, 21 April 2017
| This Sandbox is Reserved from 02/09/2015, through 05/31/2016 for use in the course "CH462: Biochemistry 2" taught by Geoffrey C. Hoops at the Butler University. This reservation includes Sandbox Reserved 1051 through Sandbox Reserved 1080. |
To get started:
More help: Help:Editing |
Diguanylate Cyclase DgcZ from Escherichia coli
| |||||||||||