This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Replication Termination Protein
From Proteopedia
(Difference between revisions)
| Line 25: | Line 25: | ||
==RTP Structure== | ==RTP Structure== | ||
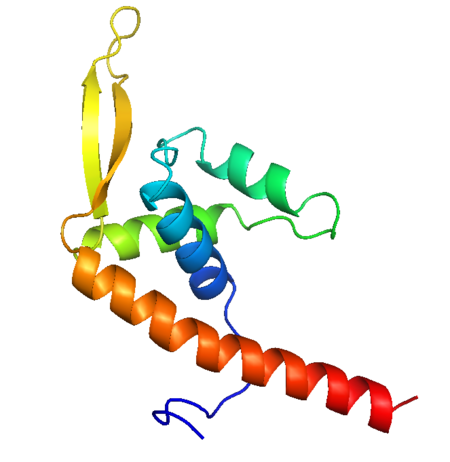
| - | RTP is a DNA binding protein from Bacillus Subtilis that uses a helix-loop-helix binding motif. In solution it shows a symmetric structure typical of the winged helix loop helix family, with an unstructured <scene name=' | + | RTP is a DNA binding protein from Bacillus Subtilis that uses a helix-loop-helix binding motif. In solution it shows a symmetric structure typical of the winged helix loop helix family, with an unstructured <scene name='44/449701/1/3'>N-terminus</scene> end, first alpha helix <scene name='Rtp_and_Tus_DNA_Binding/Alpha1/1'>(a1)</scene>, unstructured loop that is equivalent to the first beta sheet <scene name='Rtp_and_Tus_DNA_Binding/Beta1/1'>(B1)</scene>, helix loop helix structure (<scene name='Rtp_and_Tus_DNA_Binding/Alpha2/1'>a2</scene>-<scene name='Rtp_and_Tus_DNA_Binding/Alpha3/1'>a3</scene>), 2 beta sheets with a connecting loop that makes up the 'wing' structure <scene name='Rtp_and_Tus_DNA_Binding/Wing/1'>(B2 - B3)</scene> and an additional long alpha helix involved in dimerisation <scene name='Rtp_and_Tus_DNA_Binding/Alpha4/1'>(a4)</scene>.<ref>Vivian JP, Porter CJ, Wilce JA, Wilce MCJ, (2007) An asymmetric structure of the Bacillus subtilise Replication Terminator Protein in Complex with DNA. J. Mol. Bio, 370:481-491</ref> |
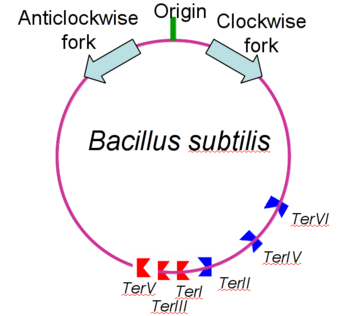
Binding of Rtp to the assymetric B portion of the Ter site changes its conformation into an assymetric 'wing-up wing-down' structure. | Binding of Rtp to the assymetric B portion of the Ter site changes its conformation into an assymetric 'wing-up wing-down' structure. | ||
Revision as of 10:19, 9 May 2017
| |||||||||||
3D structures of replication terminator protein
1bm9 – BsRTP – Bacillus subtilis
1j0r, 2dqr - BsRTP (mutant)
2dpd - BsRTP + DNA
1f4k, 2dpu, 2efw – BsRTP (mutant) + DNA
1ecr – Tus + DNA – Escherichia coli
References
- ↑ Kamada K, Horiuchi T, Ohsumi K, Shimamoto N, Morkikawa K, (1996) Structure of a replication-terminator protein complexed with DNA. Nature, 383:598-603
- ↑ Wilce et. al. (2001) Structure of the RTP−DNA complex and the mechanism of polar replication fork arrest. Nature Structural Biology, 8:206-210
- ↑ Gautam A. et.al. (2001) A single domain of the replication termination protein of Bacillus subtilis is involved in arresting both DnaB helicase and RNA polymerase. Journal of Biological Chemistry, 276:23471-23479
- ↑ Noirot P (2007). "Replication of the Bacillus subtilis chromosome". In Graumann P. Bacillus: Cellular and Molecular Biology. Caister Academic Press. ISBN 978-1-904455-12-7
- ↑ Duggin I.G. (2006) DNA Replication Fork Arrest by the Bacillus subtilis RTP–DNA Complex Involves a Mechanism that Is Independent of the Affinity of RTP–DNA Binding. Journal of Molecular Biology, 361:1-6
- ↑ Vivian JP, Porter CJ, Wilce JA, Wilce MCJ, (2007) An asymmetric structure of the Bacillus subtilise Replication Terminator Protein in Complex with DNA. J. Mol. Bio, 370:481-491
- ↑ Vivian JP, Porter CJ, Wilce JA, Wilce MCJ, (2007) An asymmetric structure of the Bacillus subtilise Replication Terminator Protein in Complex with DNA. J. Mol. Bio, 370:481-491
- ↑ Bastia D. (1995) Crystal structure of the replication terminator protein from b. subtilis at 2.6 A. Cell 80: 651-660
- ↑ Wilce et. al. (2001) Structure of the RTP−DNA complex and the mechanism of polar replication fork arrest. Nature Structural Biology, 8:206-210
- ↑ Kaplan D.L., Bastia D. (2009). Mechanisms of polar arrest of a replication fork. Molecular Microbiology 72: 279-284
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Craig Mooney, Joel L. Sussman