This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
NADH peroxidase
From Proteopedia
(Difference between revisions)
| Line 2: | Line 2: | ||
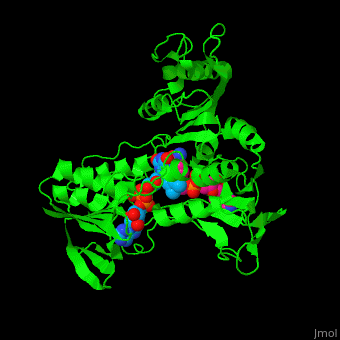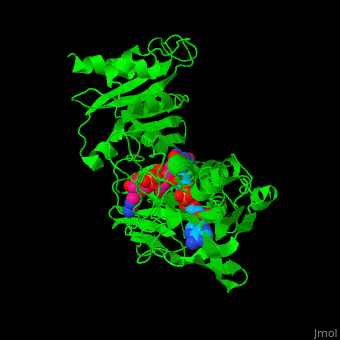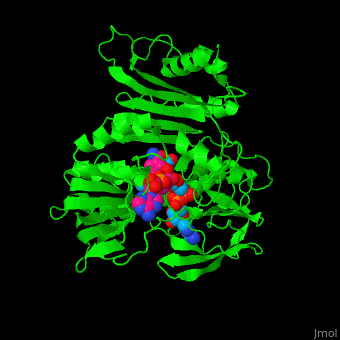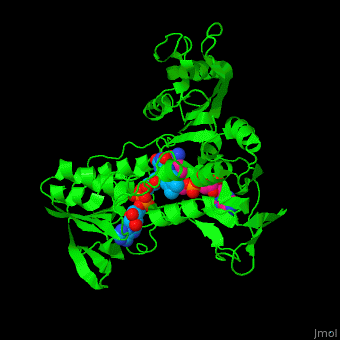
'''NADH peroxidase''' (NPO) catalyzes the conversion of NADH to NAD+ using hydrogen peroxide. See [[NAD]]. NPO eliminates the potentially toxic hydrogen peroxide and defends the cell against H<sub>2</sub>O<sub>2</sub>-mediated oxidative stress<ref>PMID:8425532</ref>. NPO provides an additional pathway for regeneration of NAD+ which is essential to the fermentative metabolism. FAD is the cofactor in this reaction. | '''NADH peroxidase''' (NPO) catalyzes the conversion of NADH to NAD+ using hydrogen peroxide. See [[NAD]]. NPO eliminates the potentially toxic hydrogen peroxide and defends the cell against H<sub>2</sub>O<sub>2</sub>-mediated oxidative stress<ref>PMID:8425532</ref>. NPO provides an additional pathway for regeneration of NAD+ which is essential to the fermentative metabolism. FAD is the cofactor in this reaction. | ||
| + | *<scene name='48/486476/Cv/4'>NAD binding site</scene>. | ||
| + | *<scene name='48/486476/Cv/8'>NAD/FAD interactions</scene>. | ||
| + | *<scene name='48/486476/Cv/7'>FAD binding site</scene>. | ||
| + | *<scene name='48/486476/Cv/9'>Whole binding site</scene>. | ||
</StructureSection> | </StructureSection> | ||
==3D structures of NADH peroxidase== | ==3D structures of NADH peroxidase== | ||
Revision as of 12:02, 10 August 2017
| |||||||||||
3D structures of NADH peroxidase
1npx, 1joa – EfNPO – Enterococcus faecalis
2npx – EfNPO + NAD
1nhp, 1nhq, 1nhr, 1nhs, 1f8w – EfNPO (mutant)
References
- ↑ Stehle T, Claiborne A, Schulz GE. NADH binding site and catalysis of NADH peroxidase. Eur J Biochem. 1993 Jan 15;211(1-2):221-6. PMID:8425532