This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
SAM-dependent methyltransferase
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
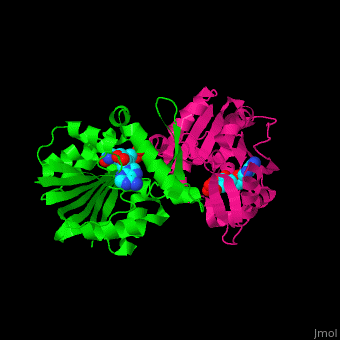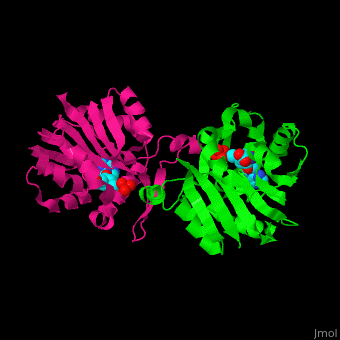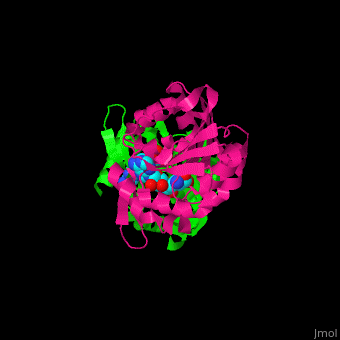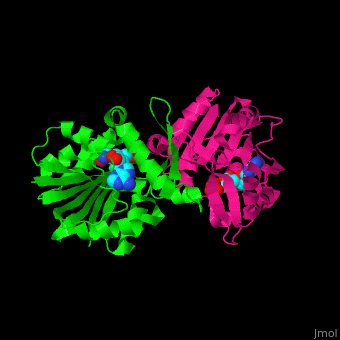
| - | <StructureSection load='' size=' | + | <StructureSection load='' size='350' side='right' scene='51/510230/Cv/3' caption='SAM-dependent methyltransferase dimer complex with S-adenosyl-L-homocysteine and sulfate [[3ou6]]'> |
== Function == | == Function == | ||
'''SAM-dependent methyltransferase''' (SDM) utilizes the methyl donor S-adenosyl-L-methionine (SAM) as a cofactor to methylate proteins, small molecules, lipids and nucleic acids. SAM forms S-adenosyl-L-homocysteine (SAH) upon demethylation. About 120 members of the SDM family have been identified. They differ in their substrate specificity and the atom targeted for methylation (N, O, C, S)<ref>PMID:23180741</ref>. | '''SAM-dependent methyltransferase''' (SDM) utilizes the methyl donor S-adenosyl-L-methionine (SAM) as a cofactor to methylate proteins, small molecules, lipids and nucleic acids. SAM forms S-adenosyl-L-homocysteine (SAH) upon demethylation. About 120 members of the SDM family have been identified. They differ in their substrate specificity and the atom targeted for methylation (N, O, C, S)<ref>PMID:23180741</ref>. | ||
| Line 37: | Line 37: | ||
**[[4kif]], [[4kig]] – ShSDM + phenylpyruvate <br /> | **[[4kif]], [[4kig]] – ShSDM + phenylpyruvate <br /> | ||
**[[4m6x]], [[4m6y]], [[4m71]] – ShSDM (mutant) + SAH + methylphenylpyruvate <br /> | **[[4m6x]], [[4m6y]], [[4m71]] – ShSDM (mutant) + SAH + methylphenylpyruvate <br /> | ||
| + | **[[5o4h]] – MmHCGC + SAM + pyridinol - Methanococcus maripaludis <br /> | ||
| + | **[[5o4n]], [[5o4j]], [[5o4m]] – MmHCGC + SAH + pyridinol <br /> | ||
*SAM-dependent methyltransferase | *SAM-dependent methyltransferase | ||
Revision as of 17:28, 17 September 2018
| |||||||||||
3D structures of SAM-dependent methyltrasferase
Updated on 17-September-2018
References
- ↑ Struck AW, Thompson ML, Wong LS, Micklefield J. S-adenosyl-methionine-dependent methyltransferases: highly versatile enzymes in biocatalysis, biosynthesis and other biotechnological applications. Chembiochem. 2012 Dec 21;13(18):2642-55. doi: 10.1002/cbic.201200556. Epub 2012, Nov 23. PMID:23180741 doi:http://dx.doi.org/10.1002/cbic.201200556
- ↑ Djordjevic S, Stock AM. Chemotaxis receptor recognition by protein methyltransferase CheR. Nat Struct Biol. 1998 Jun;5(6):446-50. PMID:9628482
- ↑ Lee JH, Bae B, Kuemin M, Circello BT, Metcalf WW, Nair SK, van der Donk WA. Characterization and structure of DhpI, a phosphonate O-methyltransferase involved in dehydrophos biosynthesis. Proc Natl Acad Sci U S A. 2010 Oct 12;107(41):17557-62. Epub 2010 Sep 27. PMID:20876132 doi:10.1073/pnas.1006848107