This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Gag polyprotein
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | |||
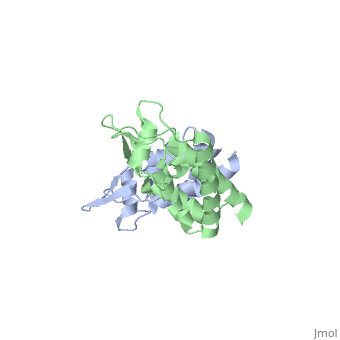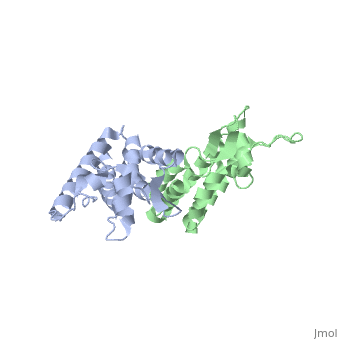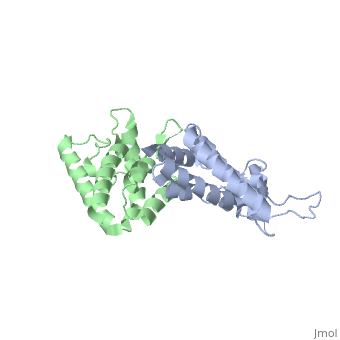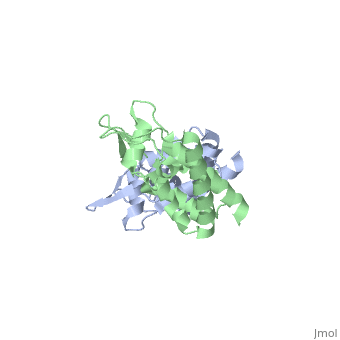
<StructureSection load='2wlv' size='350' side='right' caption='Gag polyprotein N-terminal capsid domain of HIV-2 (PDB entry [[2wlv]])' scene=''> | <StructureSection load='2wlv' size='350' side='right' caption='Gag polyprotein N-terminal capsid domain of HIV-2 (PDB entry [[2wlv]])' scene=''> | ||
| Line 17: | Line 16: | ||
==Implications== | ==Implications== | ||
HIV-1 viral particles need to form a capsid cone-like structure prior to infection of the host cell. The protealytic cleavage of the immature Gag<sup>283</sup> polyprotein results in a capsid domain. This post-translational modification is essential to the formation of the core structure. Many studies have shown that the β-hairpin formed after maturation is essential for the capsid core particle formation <ref name="gitti"/><ref name="von"/>. As a result of the β-hairpin formation, the helix 6 is displaced causing an allosteric mechanism for CpyA binding. Overall, the maturation of Gag<sup>283</sup> and formation of the mature CA protein is essential for core capsid particle creation and consequently final infection. | HIV-1 viral particles need to form a capsid cone-like structure prior to infection of the host cell. The protealytic cleavage of the immature Gag<sup>283</sup> polyprotein results in a capsid domain. This post-translational modification is essential to the formation of the core structure. Many studies have shown that the β-hairpin formed after maturation is essential for the capsid core particle formation <ref name="gitti"/><ref name="von"/>. As a result of the β-hairpin formation, the helix 6 is displaced causing an allosteric mechanism for CpyA binding. Overall, the maturation of Gag<sup>283</sup> and formation of the mature CA protein is essential for core capsid particle creation and consequently final infection. | ||
| + | |||
| + | ==3D structures of Gag polyprotein== | ||
| + | [[Gag polyprotein 3D structures]] | ||
| + | |||
</StructureSection> | </StructureSection> | ||
==3D structures of Gag polyprotein== | ==3D structures of Gag polyprotein== | ||
| Line 27: | Line 30: | ||
**[[2h3i]] – Gag residues 2-132 – HIV-1<BR /> | **[[2h3i]] – Gag residues 2-132 – HIV-1<BR /> | ||
**[[2h3f]], [[2h3i]], [[1uph]] - Gag residues 2-132 – NMR<BR /> | **[[2h3f]], [[2h3i]], [[1uph]] - Gag residues 2-132 – NMR<BR /> | ||
| + | **[[5mdg]], [[5mdf]], [[5mde]], [[5mdd]], [[5mdc]], [[5mdb]], [[5mda]], [[5md9]], [[5md8]], [[5md7]], [[5md6]], [[5md5]], [[5md4]], [[5md3]], [[5md2]], [[5md1]], [[5md0]], [[5mcz]] - Gag residues 1-221 – Cryo EM<br /> | ||
**[[1l6n]] - Gag residues 1-283 – NMR<BR /> | **[[1l6n]] - Gag residues 1-283 – NMR<BR /> | ||
**[[1gwp]] - Gag residues 132-283 – NMR<BR /> | **[[1gwp]] - Gag residues 132-283 – NMR<BR /> | ||
| Line 34: | Line 38: | ||
**[[1baj]] – Gag C terminal<BR /> | **[[1baj]] – Gag C terminal<BR /> | ||
**[[2znf]] – Gag zinc fingerlike domain - NMR<BR /> | **[[2znf]] – Gag zinc fingerlike domain - NMR<BR /> | ||
| + | **[[5teo]] - Gag residues 278-377 <br /> | ||
| + | **[[6n3j]] - Gag residues 278-377 (mutant) <br /> | ||
**[[2h3q]], [[2h3v]], [[2h3z]] - Gag residues 2-132 + phosphatidyl inositol bisphosphate - NMR<BR /> | **[[2h3q]], [[2h3v]], [[2h3z]] - Gag residues 2-132 + phosphatidyl inositol bisphosphate - NMR<BR /> | ||
**[[1mt7]], [[1mt8]] – Gag MA-CA cleavage site + protease retropepsin (mutant) <BR /> | **[[1mt7]], [[1mt8]] – Gag MA-CA cleavage site + protease retropepsin (mutant) <BR /> | ||
| Line 40: | Line 46: | ||
**[[1fgl]] – Gag residues 81-105 + cyclophilin A<BR /> | **[[1fgl]] – Gag residues 81-105 + cyclophilin A<BR /> | ||
**[[2xde]] - Gag residues 1-146 + inhibitor<BR /> | **[[2xde]] - Gag residues 1-146 + inhibitor<BR /> | ||
| - | **[[ | + | **[[4j91]], [[4j92]], [[4j93]] - Gag residues 133-278 + inhibitor<br /> |
| + | **[[6n3u]] - Gag residues 278-377 (mutant) + inhibitor<br /> | ||
**[[2x2d]] - Gag residues 133-278 + peptidyl-prolyl cis-trans isomerase A<BR /> | **[[2x2d]] - Gag residues 133-278 + peptidyl-prolyl cis-trans isomerase A<BR /> | ||
**[[2lf4]] - Gag residues 133-363 (mutant)<br /> | **[[2lf4]] - Gag residues 133-363 (mutant)<br /> | ||
| + | **[[4u0d]] - Gag residues 133-363 + Nup153 peptide<br /> | ||
| + | **[[5upw]], [[5mcy]], [[5mcx]] - Gag residues 139-351 – Cryo EM<br /> | ||
| + | **[[6ern]] - Gag residues 139-351 + ATP<br /> | ||
| + | **[[6erm]] - Gag residues 139-351 + TTP derivative<br /> | ||
| + | **[[6h09]], [[6es8]] - Gag residues 133-315 + inositol hexakisphosphate<br /> | ||
**[[2xt1]] – Gag C terminal + camelid VHH<br /> | **[[2xt1]] – Gag C terminal + camelid VHH<br /> | ||
**[[1sje]], [[1sjh]] – Gag peptide + HLA-DR1<BR /> | **[[1sje]], [[1sjh]] – Gag peptide + HLA-DR1<BR /> | ||
| Line 60: | Line 72: | ||
**[[1eoq]] - Gag C terminal – NMR<BR /> | **[[1eoq]] - Gag C terminal – NMR<BR /> | ||
**[[1a6s]] – Gag M domain (mutant) - NMR<BR /> | **[[1a6s]] – Gag M domain (mutant) - NMR<BR /> | ||
| + | **[[5a9e]] - Gag residues 84-577 – Cryo EM<br /> | ||
*Gag polyprotein from Simian immunodeficiency virus | *Gag polyprotein from Simian immunodeficiency virus | ||
| Line 69: | Line 82: | ||
**[[1u7k]], [[3bp9]] - Gag residues 215-345<BR /> | **[[1u7k]], [[3bp9]] - Gag residues 215-345<BR /> | ||
| - | **[[1u6p]] – Gag | + | **[[1u6p]] – Gag residues 479-534 + DNA <br /> |
| + | **[[6mig]] - Gag residues 683-937 + DNA<br /> | ||
*Gag polyprotein from equine infectious anemia virus | *Gag polyprotein from equine infectious anemia virus | ||
| Line 82: | Line 96: | ||
**[[4jnh]] - Gag N terminal <br /> | **[[4jnh]] - Gag N terminal <br /> | ||
| + | **[[4jmr]] - Gag N terminal + Env protein<br /> | ||
| + | **[[5m1h]], [[5m1g]] - Gag residues 300-477 - NMR<br /> | ||
}} | }} | ||
==Additional Resources== | ==Additional Resources== | ||
Revision as of 09:04, 8 July 2019
| |||||||||||
Contents |
3D structures of Gag polyprotein
Updated on 08-July-2019
Additional Resources
For additional information, see: Human Immunodeficiency Virus
Reference
- ↑ Coffin, J., S. Hughes, and H. Varmus, Retroviruses. 1997: Cold Spring Harbor Laboratory Press.
- ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedsource - ↑ 3.0 3.1 Gitti RK, Lee BM, Walker J, Summers MF, Yoo S, Sundquist WI. Structure of the amino-terminal core domain of the HIV-1 capsid protein. Science. 1996 Jul 12;273(5272):231-5. PMID:8662505
- ↑ 4.0 4.1 von Schwedler UK, Stemmler TL, Klishko VY, Li S, Albertine KH, Davis DR, Sundquist WI. Proteolytic refolding of the HIV-1 capsid protein amino-terminus facilitates viral core assembly. EMBO J. 1998 Mar 16;17(6):1555-68. PMID:9501077 doi:10.1093/emboj/17.6.1555
- ↑ Braaten D, Franke EK, Luban J. Cyclophilin A is required for an early step in the life cycle of human immunodeficiency virus type 1 before the initiation of reverse transcription. J Virol. 1996 Jun;70(6):3551-60. PMID:8648689
- ↑ Thali M, Bukovsky A, Kondo E, Rosenwirth B, Walsh CT, Sodroski J, Gottlinger HG. Functional association of cyclophilin A with HIV-1 virions. Nature. 1994 Nov 24;372(6504):363-5. PMID:7969495 doi:http://dx.doi.org/10.1038/372363a0
- ↑ Ackerson B, Rey O, Canon J, Krogstad P. Cells with high cyclophilin A content support replication of human immunodeficiency virus type 1 Gag mutants with decreased ability to incorporate cyclophilin A. J Virol. 1998 Jan;72(1):303-8. PMID:9420228
Team from University of Missouri, Columbia, MO
- Students: Zheng Wang, Allison Tegge, Xin Deng
- Advisors: Jianlin Cheng, PhD, Department of Computer Science, Informatics Institute, the Life Science Center, Interdisciplinary Plant Group, University of Missouri, Columbia
- Mentor: Chun Tang, PhD, Department of Biochemistry, University of Missouri, Columbia
NMR Equipment and the Authors
Created by Allison Tegge and David Canner