This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Alcohol dehydrogenase
From Proteopedia
(Difference between revisions)
| Line 23: | Line 23: | ||
<br/> | <br/> | ||
{{Clear}} | {{Clear}} | ||
| - | |||
| - | ==Structure== | ||
| - | |||
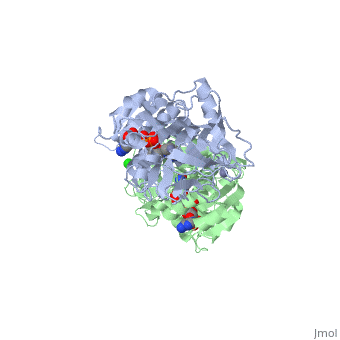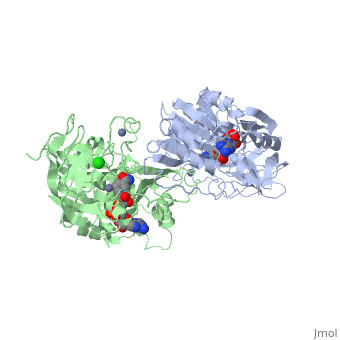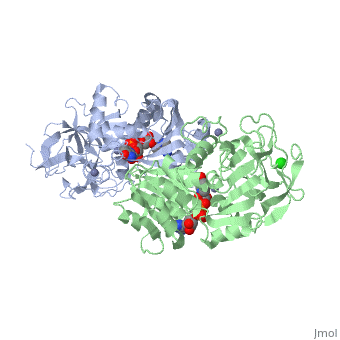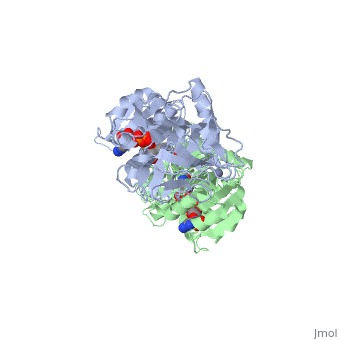
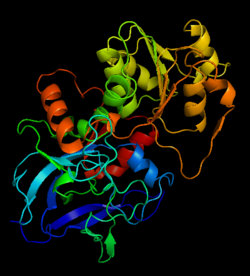
| - | The initial scene (<scene name='Birrer_Sandbox_2/Whole_adh_molecule/3'>Domains of ADH</scene>) shows an overview of the molecule, allowing for a general look at the tertiary structure of alcohol dehydrogenase (it is complexed with Cl, Pyz, NAD, and Zn). A second scene (<scene name='Birrer_Sandbox_2/Close_look_at_ligand/2'>Closer Look at Subunit</scene>) shows a close view of the ligand within each subunit. Labels have been placed on NAD, CL, and Zn to clearly establish the structure. | ||
| - | |||
| - | |||
| - | Within alcohol dehydrogenase, <scene name='Birrer_Sandbox_2/The_active_site/1'>the active</scene> site of alcohol dehydrogenase has three important residues, Phe 93, Leu 57, and Leu 116. These three residues work together to bind to the alcohol substrate.<ref>''Protein: Alcohol Dehydrogenase''. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://web.archive.org/web/20080307193453/http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm></ref> | ||
| - | |||
| - | |||
| - | Zn plays an important role in the catalysis. It funtions by electrostatically stabilizing the oxygen in alcohol during the reaction, which causes the alcohol to be more acidic. At the <scene name='Birrer_Sandbox_2/Zinc_binding_site/1'>Zinc Binding Site</scene>, Zinc coordinates with Cys 146, Cys 174, and His 67.<ref>''Protein: Alcohol Dehydrogenase''. The College of Saint Benedict and Saint John's University. 1 March 2010 <http://web.archive.org/web/20080307193453/http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm></ref> | ||
| - | |||
| - | |||
| - | NAD functions as a cosubstrate in the dehydration. NAD binds to numerous residues in a series of beta-alpha-beta folds. <scene name='Birrer_Sandbox_2/Nad_binding_site/1'>NAD Binding Region</scene> shows the domain where NAD binds, and many of the residues with which it interacts are selected. | ||
| - | <ref>''Protein: Alcohol Dehydrogenase''. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://web.archive.org/web/20080307193453/http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm</ref> | ||
| - | |||
| - | |||
| - | Alcohol dehydrogenase exists as a dimer with a zinc molecule complexed in each of the subunits. It has a SCOP catagory of an alpha and beta protein. At the N-terminal, there is a domain that is all beta; however, the C-Terminal domain is alpha and beta, so the catagory is alpha and beta. The C-Terminal core has 3 layers of alpha/beta/alpha and parallel beta sheets of 6 strands.<ref>''Protein: Alcohol dehydrogenase from Human (Homo sapiens), different isozymes''. SCOP. 2009. 1 March 2010 < http://web.archive.org/web/20060914235939/http://scop.berkeley.edu/data/scop.b.d.c.b.b.c.html></ref> | ||
| - | |||
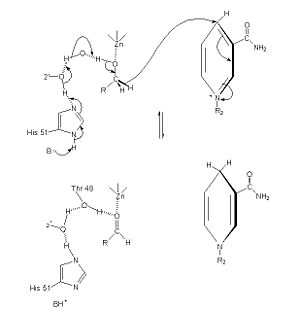
==Reaction and Mechanism== | ==Reaction and Mechanism== | ||
Revision as of 11:24, 30 May 2018
| |||||||||||
Additional Resources
For additional information, see: Carbohydrate Metabolism
3D Structures of Alcohol dehydrogenase
Updated on 30-May-2018
References
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://web.archive.org/web/20080307193453/http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://web.archive.org/web/20080307193453/http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Bille V, Remacle J. Simple-kinetic descriptions of alcohol dehydrogenase after immobilization on tresyl-chloride-activated agarose. Eur J Biochem. 1986 Oct 15;160(2):343-8. PMID:3769934
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Blomstrand R, Ostling-Wintzell H, Lof A, McMartin K, Tolf BR, Hedstrom KG. Pyrazoles as inhibitors of alcohol oxidation and as important tools in alcohol research: an approach to therapy against methanol poisoning. Proc Natl Acad Sci U S A. 1979 Jul;76(7):3499-503. PMID:115004
- ↑ Alcohol Dehydrogenase. Worthington Biochemical Corporation . 31 March 2010 < http://http://www.worthington-biochem.com/ADH/default.html>
- ↑ Alcohol Dehydrogenase.Worthington Biochemical Corporation . 31 March 2010 < http://http://www.worthington-biochem.com/ADH/default.html>
- ↑ Goihberg E, Dym O, Tel-Or S, Levin I, Peretz M, Burstein Y. A single proline substitution is critical for the thermostabilization of Clostridium beijerinckii alcohol dehydrogenase. Proteins. 2007 Jan 1;66(1):196-204. PMID:17063493 doi:10.1002/prot.21170
- ↑ Goihberg E, Dym O, Tel-Or S, Shimon L, Frolow F, Peretz M, Burstein Y. Thermal stabilization of the protozoan Entamoeba histolytica alcohol dehydrogenase by a single proline substitution. Proteins. 2008 Feb 7;. PMID:18260103 doi:10.1002/prot.21946
- ↑ Goihberg E, Peretz M, Tel-Or S, Dym O, Shimon L, Frolow F, Burstein Y. Biochemical and Structural Properties of Chimeras Constructed by Exchange of Cofactor-Binding Domains in Alcohol Dehydrogenases from Thermophilic and Mesophilic Microorganisms. Biochemistry. 2010 Feb 9. PMID:20102159 doi:10.1021/bi901730x
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, David Canner, Joel L. Sussman, David Birrer