This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Pho4
From Proteopedia
| Line 9: | Line 9: | ||
The pho4 protein takes part in transcription of several yeast genes related to phosphate metabolism: | The pho4 protein takes part in transcription of several yeast genes related to phosphate metabolism: | ||
| - | [[PH05]] encoding repressible acid phosphatase, | + | '''[[PH05]] encoding repressible acid phosphatase,''' the expression of Pho5 in Saccharomyces cerevisiae is derepressed by a specific level of inorganic phosphate (Pi), and depressed when the Pi concentrate in the medium is lowered, the regulatory system of the Pi signals coded by PHO4 PHO2 , PHO80, and PHO81 genes.<ref>PMID:2671650</ref> |
| - | [[PH08]] encoding alkaline phosphatase<ref>PMID:2671650</ref> | + | '''[[PH08]] encoding alkaline phosphatase'''<ref>PMID:2671650</ref> |
| - | [[PH081 ]] a regulatory gene<ref>PMID:2671650</ref> | + | '''[[PH081 ]] a regulatory gene''' regulated by PHO system wuth the use of Pi<ref>PMID:2671650</ref> |
| - | [[PH084]] phosphate permease gene<ref>PMID:3902805</ref> | + | '''[[PH084]] phosphate permease gene'''<ref>PMID:3902805</ref> |
other than that, the Pho4 protein seems to contain a coding region for non translated mitochondrial proteins<ref>PMID:2183025</ref> | other than that, the Pho4 protein seems to contain a coding region for non translated mitochondrial proteins<ref>PMID:2183025</ref> | ||
| Line 23: | Line 23: | ||
==sturcture== | ==sturcture== | ||
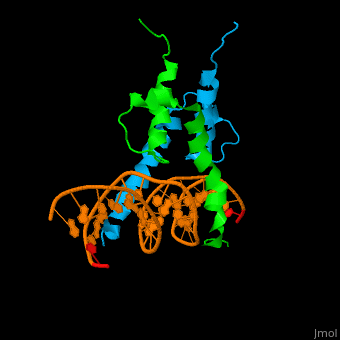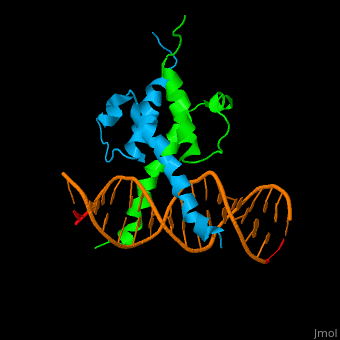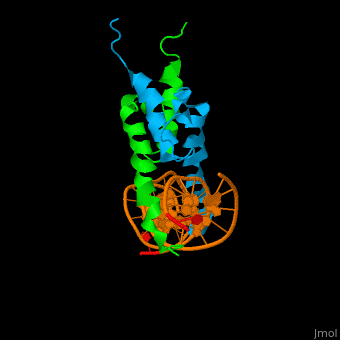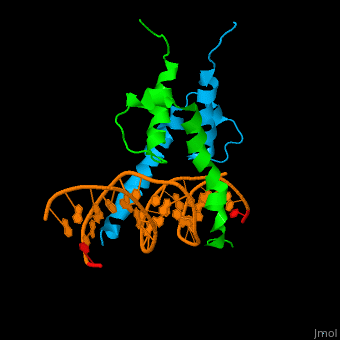
| - | The structure of the Pho4 protein is a basic Helix-loop-helix(bHLH) , The linker sequence between Helix1(H1) and Helix2(H2) contains several residues: Asn, Glu, Pro and Ser<ref>PMID:3775366</ref>. The DNA binding site is a homodimer and two monomers in a left‐handed four‐helix bundle fold (identical to bHLH/Zip and bHLH proteins fold) . | + | The structure of the Pho4 protein is a basic Helix-loop-helix(bHLH), The linker sequence between Helix1(H1) and Helix2(H2) contains several residues: Asn, Glu, Pro and Ser<ref>PMID:3775366</ref>. The DNA binding site is a homodimer and two monomers in a left‐handed four‐helix bundle fold (identical to bHLH/Zip and bHLH proteins fold) . |
*loop structure | *loop structure | ||
Revision as of 14:47, 19 March 2018
| |||||||||||
sturcture
The structure of the Pho4 protein is a basic Helix-loop-helix(bHLH), The linker sequence between Helix1(H1) and Helix2(H2) contains several residues: Asn, Glu, Pro and Ser[8]. The DNA binding site is a homodimer and two monomers in a left‐handed four‐helix bundle fold (identical to bHLH/Zip and bHLH proteins fold) .
- loop structure
Pho4 loops (conserved structure?) are long but compact and forms a short alpha-helical structure [9]. The PHO4 loop contains a Trp residue that faces the aromatic rings of Tyr52 and His55 of helix H2, and Pro28 of H2 and H1.an aromatic cluster forms a cap structure is observed.[10]
- DNA binding
As The Pho4 binding region binds specifically to CACGTG sequence by a specific subset of the basic region of the bHLH motif[11]
3D Structures of lactoferrin
Updated on 19-March-2018
1a0a – yPho4 DNA-binding domain + DNA - yeast
3w3x – yPho4 peptide + importin subunit b-3
References
- ↑ Berben G, Legrain M, Gilliquet V, Hilger F. The yeast regulatory gene PHO4 encodes a helix-loop-helix motif. Yeast. 1990 Sep-Oct;6(5):451-4. PMID:2220078 doi:http://dx.doi.org/10.1002/yea.320060510
- ↑ Berben G, Legrain M, Gilliquet V, Hilger F. The yeast regulatory gene PHO4 encodes a helix-loop-helix motif. Yeast. 1990 Sep-Oct;6(5):451-4. PMID:2220078 doi:http://dx.doi.org/10.1002/yea.320060510
- ↑ Yoshida K, Ogawa N, Oshima Y. Function of the PHO regulatory genes for repressible acid phosphatase synthesis in Saccharomyces cerevisiae. Mol Gen Genet. 1989 May;217(1):40-6. PMID:2671650
- ↑ Yoshida K, Ogawa N, Oshima Y. Function of the PHO regulatory genes for repressible acid phosphatase synthesis in Saccharomyces cerevisiae. Mol Gen Genet. 1989 May;217(1):40-6. PMID:2671650
- ↑ Yoshida K, Ogawa N, Oshima Y. Function of the PHO regulatory genes for repressible acid phosphatase synthesis in Saccharomyces cerevisiae. Mol Gen Genet. 1989 May;217(1):40-6. PMID:2671650
- ↑ Tamai Y, Toh-e A, Oshima Y. Regulation of inorganic phosphate transport systems in Saccharomyces cerevisiae. J Bacteriol. 1985 Nov;164(2):964-8. PMID:3902805
- ↑ Ogawa N, Oshima Y. Functional domains of a positive regulatory protein, PHO4, for transcriptional control of the phosphatase regulon in Saccharomyces cerevisiae. Mol Cell Biol. 1990 May;10(5):2224-36. PMID:2183025
- ↑ Leszczynski JF, Rose GD. Loops in globular proteins: a novel category of secondary structure. Science. 1986 Nov 14;234(4778):849-55. PMID:3775366
- ↑ Schmeckpeper BJ, Adams JM, Harris AW. Detection of a possible precursor of immunoglobulin light chain in MOPC 41 A plasmacytoma cells. FEBS Lett. 1975 Apr 15;53(1):95-8. PMID:1170095
- ↑ Shimizu T, Toumoto A, Ihara K, Shimizu M, Kyogoku Y, Ogawa N, Oshima Y, Hakoshima T. Crystal structure of PHO4 bHLH domain-DNA complex: flanking base recognition. EMBO J. 1997 Aug 1;16(15):4689-97. PMID:9303313 doi:10.1093/emboj/16.15.4689
- ↑ Fisher F, Jayaraman PS, Goding CR. C-myc and the yeast transcription factor PHO4 share a common CACGTG-binding motif. Oncogene. 1991 Jul;6(7):1099-104. PMID:1861859
Proteopedia Page Contributors and Editors (what is this?)
Inbar Yaffe, Michal Harel, Alexander Berchansky, Joel L. Sussman