This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Kyle Burton/Sandbox1
From Proteopedia
| Line 11: | Line 11: | ||
== Structure == | == Structure == | ||
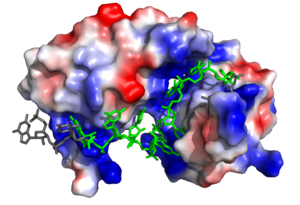
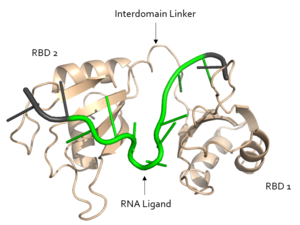
[[Image:Sex Lethal Protein Structural Overview with Labels.png|300px|right|thumb| '''Figure 2.''' Structural overview of Sxl. RNA ligand colored in green is recognized and bound, while RNA ligand colored in grey is not bound. Image created in PyMol. Structure shown is [https://www.rcsb.org/structure/1b7f PDB:1b7f].]] | [[Image:Sex Lethal Protein Structural Overview with Labels.png|300px|right|thumb| '''Figure 2.''' Structural overview of Sxl. RNA ligand colored in green is recognized and bound, while RNA ligand colored in grey is not bound. Image created in PyMol. Structure shown is [https://www.rcsb.org/structure/1b7f PDB:1b7f].]] | ||
| - | Sxl is composed of two asymmetric RNA binding domains (RBD1 and RBD2) which recognize a poly-uridine site in the pre-mRNA transcript<ref name="Handa"/>. Each RBD is comprised of two alpha helices and one antiparallel four-stranded β sheet<ref name="Handa"/>. The β sheets face each other, lining the electropositive V-shaped cleft<ref name="Handa"/>. The inter-domain linker forms a distorted 310 helix which helps to form the V-shaped cleft into which the pre-mRNA sequence binds<ref name="Handa"/><ref name="Black">doi: 10.1146/annurev.biochem.72.121801.161720</ref>. Sxl binds to UGUUUUUUU sequence of GUUGUUUUUUUU in tra. RBD1 binds U6-U11 and RBD2 binds U3, G4, and U5. Although the two RBDs do not interact with each other, this nine-ribonucleotide sequence must be recognized continuously to prevent U2AF from binding at the 3’ splice site. The binding of Sxl to the pre-mRNA occurs in an electropositive pocket due to extensive interactions with the RNA phosphate backbone and negatively charged residues. Since Sxl binds primarily with the phosphate backbone, the protein residues are not highly conserved. | + | Sxl is composed of two asymmetric RNA binding domains (RBD1 and RBD2) which recognize a poly-uridine site in the pre-mRNA transcript<ref name="Handa"/>. Each RBD is comprised of two alpha helices and one antiparallel four-stranded β sheet<ref name="Handa"/>. The β sheets face each other, lining the electropositive V-shaped cleft<ref name="Handa"/>. The inter-domain linker forms a distorted 310 helix which helps to form the V-shaped cleft into which the pre-mRNA sequence binds<ref name="Handa"/><ref name="Black">doi: 10.1146/annurev.biochem.72.121801.161720</ref>. Sxl binds to UGUUUUUUU sequence of GUUGUUUUUUUU in tra. RBD1 binds U6-U11 and RBD2 binds U3, G4, and U5. Although the two RBDs do not interact with each other, this nine-ribonucleotide sequence must be recognized continuously to prevent U2AF from binding at the 3’ splice site<ref name="Handa"/>. The binding of Sxl to the pre-mRNA occurs in an electropositive pocket due to extensive interactions with the RNA phosphate backbone and negatively charged residues<ref name="Handa"/>. Since Sxl binds primarily with the phosphate backbone, the protein residues are not highly conserved. |
=== Alternative Splicing Pathways === | === Alternative Splicing Pathways === | ||
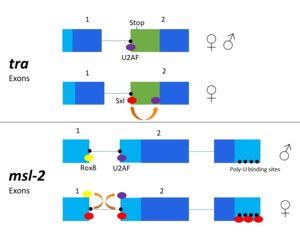
[[Image:Sxl mechanism alternativesplicing figure version2.jpg|300px|left|thumb| '''Figure 3.''' 2-dimensional representation of alternative splicing repression by Sxl on the ''tra'' and ''msl-2'' genes.]] | [[Image:Sxl mechanism alternativesplicing figure version2.jpg|300px|left|thumb| '''Figure 3.''' 2-dimensional representation of alternative splicing repression by Sxl on the ''tra'' and ''msl-2'' genes.]] | ||
| - | In alternative splicing of the ''tra'' gene, Sxl binds at the 3' poly-uridine site. This causes U2AF to bind downstream and the spliceosome transcribes the following exon<ref name="Penalva"/>. | + | In alternative splicing of the ''tra'' gene, Sxl binds at the 3' poly-uridine site. This causes U2AF to bind downstream and the spliceosome transcribes the following exon<ref name="Penalva"/>. In the absence of Sxl, the normal gene for male development is transcribed. The exon contains a stop codon which results in a truncated, non-functional protein<ref name="Black"/>. In the presence of Sxl, this exon is spliced, so the stop codon is skipped<ref name="Black"/>. This enables translation of an active ''tra'' protein<ref name="Black"/>. |
=== Structural Highlights === | === Structural Highlights === | ||
Revision as of 14:54, 29 March 2018
Contents |
Sex-Lethal Protein
| |||||||||||
Additional Reading
For more information on the U2AF splicing factor.
Relevance
As Sxl functions as a splicing repressor, it may give insight into the effects of varying mechanisms of alternate splicing both in flies and other species. Sxl may also lead to understanding of human alternative splicing factors. As an RNA binding protein, research regarding Sxl may contribute to the understanding of enzymes with RNA recognition motifs.
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 Handa N, Nureki O, Kurimoto K, Kim I, Sakamoto H, Shimura Y, Muto Y, Yokoyama S. Structural basis for recognition of the tra mRNA precursor by the Sex-lethal protein. Nature. 1999 Apr 15;398(6728):579-85. PMID:10217141 doi:10.1038/19242
- ↑ 2.0 2.1 Penalva LO, Sanchez L. RNA binding protein sex-lethal (Sxl) and control of Drosophila sex determination and dosage compensation. Microbiol Mol Biol Rev. 2003 Sep;67(3):343-59, table of contents. PMID:12966139
- ↑ 3.0 3.1 3.2 3.3 Black DL. Mechanisms of alternative pre-messenger RNA splicing. Annu Rev Biochem. 2003;72:291-336. doi: 10.1146/annurev.biochem.72.121801.161720., Epub 2003 Feb 27. PMID:12626338 doi:http://dx.doi.org/10.1146/annurev.biochem.72.121801.161720