This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Kyle Burton/Sandbox1
From Proteopedia
| Line 6: | Line 6: | ||
== Significance == | == Significance == | ||
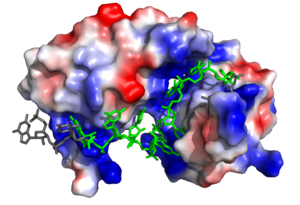
| - | [[Image:Sex lethal protein electrostatic surface representation.png|300px|right|thumb| '''Figure 1.''' Three-dimensional representation of Sex-lethal protein showing the | + | [[Image:Sex lethal protein electrostatic surface representation.png|300px|right|thumb| '''Figure 1.''' Three-dimensional representation of Sex-lethal protein showing the electropositive binding pocket and the bound RNA ligand. Pre-mRNA residues binding to Sxl shown in green, non-binding residues shown in grey. Structure shown is [https://www.rcsb.org/structure/1b7f PDB:1b7f]. Image created in PyMol.]] |
The Sxl RNA splicing targets encode for the transformer (''tra'') and the male-sex lethal (''msl-2'') proteins. Tra is a splicing activator for the female developmental pathway, and Msl-2 modulates [https://en.wikipedia.org/wiki/X_chromosome X chromosome] application in male fruit flies. The mechanism for how Sxl targets these pathways differs slightly. In both mechanisms, Sxl occupies the 3' splice site and prevents [https://en.wikipedia.org/wiki/U2AF2 U2AF] from binding. This causes the U2AF splicing factor to bind at a downstream splice site encoding proteins in the female developmental pathway. In Msl-2 targeting, Sxl also blocks the binding of another regulatory splicing factor, TIA-1, and the [https://en.wikipedia.org/wiki/SnRNP U1 snRNP] at the 5’ splice site. Sxl can also control its own splicing pattern to conserve female expression. It does so by binding to [https://en.wikipedia.org/wiki/Exon Exon] 3 of its own RNA and creating an RNP complex to eliminate this exon. After removal of Exon 3, Sxl becomes active and female expression is maintained. | The Sxl RNA splicing targets encode for the transformer (''tra'') and the male-sex lethal (''msl-2'') proteins. Tra is a splicing activator for the female developmental pathway, and Msl-2 modulates [https://en.wikipedia.org/wiki/X_chromosome X chromosome] application in male fruit flies. The mechanism for how Sxl targets these pathways differs slightly. In both mechanisms, Sxl occupies the 3' splice site and prevents [https://en.wikipedia.org/wiki/U2AF2 U2AF] from binding. This causes the U2AF splicing factor to bind at a downstream splice site encoding proteins in the female developmental pathway. In Msl-2 targeting, Sxl also blocks the binding of another regulatory splicing factor, TIA-1, and the [https://en.wikipedia.org/wiki/SnRNP U1 snRNP] at the 5’ splice site. Sxl can also control its own splicing pattern to conserve female expression. It does so by binding to [https://en.wikipedia.org/wiki/Exon Exon] 3 of its own RNA and creating an RNP complex to eliminate this exon. After removal of Exon 3, Sxl becomes active and female expression is maintained. | ||
== Structure == | == Structure == | ||
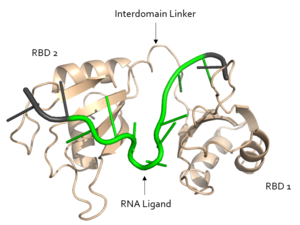
[[Image:Sex Lethal Protein Structural Overview with Labels.png|300px|right|thumb| '''Figure 2.''' Structural overview of Sxl. RNA ligand colored in green is recognized and bound, while RNA ligand colored in grey is not bound. Image created in PyMol. Structure shown is [https://www.rcsb.org/structure/1b7f PDB:1b7f].]] | [[Image:Sex Lethal Protein Structural Overview with Labels.png|300px|right|thumb| '''Figure 2.''' Structural overview of Sxl. RNA ligand colored in green is recognized and bound, while RNA ligand colored in grey is not bound. Image created in PyMol. Structure shown is [https://www.rcsb.org/structure/1b7f PDB:1b7f].]] | ||
| - | Sxl is composed of two asymmetric RNA binding domains (RBD1 and RBD2) which recognize a poly-uridine site in the pre-mRNA transcript<ref name="Handa"/>. Each RBD is comprised of two alpha helices and one antiparallel four-stranded β sheet<ref name="Handa"/>. The β sheets face each other, lining the electropositive V-shaped cleft<ref name="Handa"/>. The inter-domain linker forms a distorted 3<sub>10</sub> helix which helps form the V-shaped cleft into which the pre-mRNA sequence binds<ref name="Handa"/><ref name="Black">doi: 10.1146/annurev.biochem.72.121801.161720</ref>. Sxl binds to UGUUUUUUU sequence of GUUGUUUUUUUU in tra. RBD1 binds U6-U11 and RBD2 binds U3, G4, and U5. Although the two RBDs do not interact with each other, this nine-ribonucleotide sequence must be recognized continuously to | + | Sxl is composed of two asymmetric RNA binding domains (RBD1 and RBD2) which recognize a poly-uridine site in the pre-mRNA transcript<ref name="Handa"/>. Each RBD is comprised of two alpha helices and one antiparallel four-stranded β sheet<ref name="Handa"/>. The β sheets face each other, lining the electropositive V-shaped cleft<ref name="Handa"/>. The inter-domain linker forms a distorted 3<sub>10</sub> helix which helps form the V-shaped cleft into which the pre-mRNA sequence binds<ref name="Handa"/><ref name="Black">doi: 10.1146/annurev.biochem.72.121801.161720</ref>. Sxl binds to UGUUUUUUU sequence of GUUGUUUUUUUU in the ''tra'' pre-mRNA. RBD1 binds U6-U11 and RBD2 binds U3, G4, and U5. Although the two RBDs do not interact with each other, this nine-ribonucleotide sequence must be recognized continuously to allow Sxl to bind, preventing U2AF from binding at the 3’ splice site<ref name="Handa"/>. The binding of Sxl to the pre-mRNA occurs in an electropositive pocket due to extensive interactions with the RNA phosphate backbone and negatively charged residues<ref name="Handa"/>. Since Sxl binds primarily with the phosphate backbone, the protein residues are not highly conserved. |
=== Alternative Splicing Pathways === | === Alternative Splicing Pathways === | ||
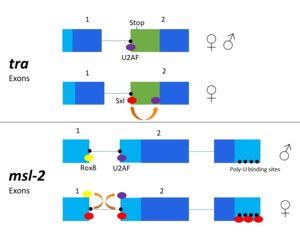
[[Image:Sxl mechanism alternativesplicing figure version2.jpg|300px|left|thumb| '''Figure 3.''' 2-dimensional representation of alternative splicing repression by Sxl on the ''tra'' and ''msl-2'' genes.]] | [[Image:Sxl mechanism alternativesplicing figure version2.jpg|300px|left|thumb| '''Figure 3.''' 2-dimensional representation of alternative splicing repression by Sxl on the ''tra'' and ''msl-2'' genes.]] | ||
| - | The alternative splicing pathways of Sxl differ, but both involve repression at the 3' splice site. The ''tra'' expression pathway only involves the 3' splice site, while the ''msl-2'' pathway involves both the 3' splice site and the 5' splice site. Both mechanisms cause U2AF binding downstream (Fig. 2)<ref name="Black"/>. U2AF is a more general splicing factor than Sxl, | + | The alternative splicing pathways of Sxl differ, but both involve repression at the 3' splice site. The ''tra'' expression pathway only involves the 3' splice site, while the ''msl-2'' pathway involves both the 3' splice site and the 5' splice site. Both mechanisms cause U2AF binding downstream (Fig. 2)<ref name="Black"/>. U2AF is a more general splicing factor than Sxl, and prefers cytidine-containing poly-uridine pre-mRNA sequences, so Sxl binds to the guanosine-containing pre-mRNA with a 10<sup>4</sup>-fold greater affinity. |
==== Autoregulation ==== | ==== Autoregulation ==== | ||
| - | Sxl is capable of autoregulation of its expression<ref name="Black"/>. The Sxl gene is transcribed in male flies, but the inclusion of exon 3 results in a premature stop codon, producing an inactive, truncated protein. The same Sxl promoter is active in female flies, but an additional (briefly active) Sxl promoter produces a transcript with exon 3 removed, | + | Sxl is capable of autoregulation of its expression<ref name="Black"/>. The Sxl gene is transcribed in male flies, but the inclusion of exon 3 results in a premature stop codon, producing an inactive, truncated protein. The same Sxl promoter is active in female flies, but an additional (briefly active) Sxl promoter produces a transcript with exon 3 removed, resulting in an active Sxl protein which will initiate other female-specific splicing cascades<ref name="Black"/>. |
==== ''Tra'' ==== | ==== ''Tra'' ==== | ||
| Line 29: | Line 29: | ||
=== Structural Basis for Recognition of Poly-U Sequences === | === Structural Basis for Recognition of Poly-U Sequences === | ||
| - | The | + | The structural interactions with regards to the targeting of the 5' splice site and of its own mRNA transcript are much less understood than the competition of Sxl with U2AF at the 3' splice site. All the RNA-protein interactions described here refer to ''tra'' pre-mRNA-Sxl interactions. |
| - | + | The <scene name='78/783145/Arg_252_interaction_with_u3_g4/6'>Arg-252 interaction with U3 and G4</scene> is crucial to pre-mRNA binding; a mutation of Arg252 to alanine eliminated the ability of Sxl to bind RNA<ref name="Handa"/>. | |
| - | The | + | |
| + | The ligand pre-mRNA sequence forms a characteristic <scene name='78/783145/U5_u6_u7_loop/2'>loop</scene> at U5, U6, and U7. This interaction is stabilized by π stacking between the G4 and U5 nucleotides and residues <scene name='78/783145/Aromatic_stacking/3'>Tyr 214 and Phe 256</scene>, respectively<ref name="Handa"/>. The nucleobases are exposed to residues on Sxl due to the 2’ endo conformation of all the nucleotides except for U8, which maintains a 3’ endo conformation<ref name="Handa"/>. | ||
| + | |||
| + | The U6 residue is recognized as part of the RNA <scene name='78/783145/U5_u6_u7_loop/2'>loop</scene> by <scene name='78/783145/Molecule_base_origin/4'>Arg195</scene><ref name="Handa"/>. The Arg195 amide hydrogen-bonds to the O2' of U6 and the U6 N3H hydrogen bonds to the Arg195 carbonyl oxygen<ref name="Handa"/>. | ||
| Line 38: | Line 41: | ||
| - | <scene name='78/783145/Arg_252_interaction_with_u3_g4/6'>Arg-252 interaction with U3 and G4</scene> | ||
<scene name='78/783145/Arg_258_interaction_w_u9_u10/3'>Arg-258 interaction with U9 and U10</scene> | <scene name='78/783145/Arg_258_interaction_w_u9_u10/3'>Arg-258 interaction with U9 and U10</scene> | ||
| Line 45: | Line 47: | ||
<scene name='78/783145/R155_intxn_with_u11/3'>R155 interaction with U11</scene> | <scene name='78/783145/R155_intxn_with_u11/3'>R155 interaction with U11</scene> | ||
| - | |||
| - | <scene name='78/783145/Aromatic_stacking/3'>Aromatic stacking between G4/Tyr-214 and U5/Phe-256</scene> | ||
<scene name='78/783145/U8_with_s165_and_y166/1'>U8 interaction with Ser-165 and Tyr-166</scene> | <scene name='78/783145/U8_with_s165_and_y166/1'>U8 interaction with Ser-165 and Tyr-166</scene> | ||
| - | <scene name='78/783145/U5_u6_u7_loop/2'>U5-U7 Loop</scene> | ||
<scene name='78/783145/U7_u8_stacking/1'>U7-U8 Stacking</scene> | <scene name='78/783145/U7_u8_stacking/1'>U7-U8 Stacking</scene> | ||
| Line 56: | Line 55: | ||
<scene name='78/783145/U9_with_interdomain_linker/1'>U9 with Interdomain Linker</scene> | <scene name='78/783145/U9_with_interdomain_linker/1'>U9 with Interdomain Linker</scene> | ||
| - | + | ||
Revision as of 16:28, 29 March 2018
Contents |
Sex-Lethal Protein
| |||||||||||
Additional Reading
For more information on the U2AF splicing factor.
Relevance
As Sxl functions as a splicing repressor, it may give insight into the effects of varying mechanisms of alternate splicing both in flies and other species. Sxl may also lead to understanding of human alternative splicing factors. As an RNA binding protein, research regarding Sxl may contribute to the understanding of enzymes with RNA recognition motifs.
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 Handa N, Nureki O, Kurimoto K, Kim I, Sakamoto H, Shimura Y, Muto Y, Yokoyama S. Structural basis for recognition of the tra mRNA precursor by the Sex-lethal protein. Nature. 1999 Apr 15;398(6728):579-85. PMID:10217141 doi:10.1038/19242
- ↑ 2.0 2.1 Penalva LO, Sanchez L. RNA binding protein sex-lethal (Sxl) and control of Drosophila sex determination and dosage compensation. Microbiol Mol Biol Rev. 2003 Sep;67(3):343-59, table of contents. PMID:12966139
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 3.6 Black DL. Mechanisms of alternative pre-messenger RNA splicing. Annu Rev Biochem. 2003;72:291-336. doi: 10.1146/annurev.biochem.72.121801.161720., Epub 2003 Feb 27. PMID:12626338 doi:http://dx.doi.org/10.1146/annurev.biochem.72.121801.161720
- ↑ doi: https://dx.doi.org/10.1128/mmbr.67.3.343-359.2003