This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Poly(A) binding protein
From Proteopedia
(Difference between revisions)
| Line 22: | Line 22: | ||
====Adenosine Stabilization Interaction Patterns==== | ====Adenosine Stabilization Interaction Patterns==== | ||
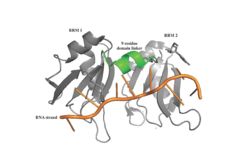
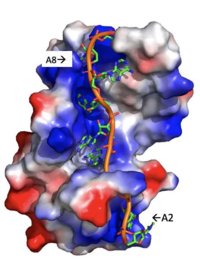
| - | Specifically, there are several significant interaction patterns that stabilize adenosine recognition. RRM 1 and 2 makes significant interactions with the adenosine backbone, shown in Figure 3. Additionally, the adenosine stabilizes itself within the binding by intramolecular stacking interactions between adenosines. Through the extensive <scene name='78/781949/Lys_104_asp_105/1'>interactions with adenosine 2</scene>, the RRM specifies the position of adenosine 2, allowing it to make strong intramolecular stacking interactions with adenosine 1. As a result, adenosine 1 requires less contact with the RRM, as it is mostly stabilized by adenosine 2. Furthermore, some adenosines like adenosine 3 and adenosine 6 are stabilized by being sandwiched between aromatic and alipathic side chains. <scene name='78/781947/Interactions_with_a3/1'>Adenosine-3 sandwiching</scene> occurs between aromatic and alipathic side chains and is specified by Lysine 104, and <scene name='78/781947/Residues_interacting_with_a6/1'>Adenosine-6 sandwiching</scene> occurs similarly, but it is specified doubly by two residues, Trp-86 and Gln-88. <ref> Deo, Rahul C, et al. “Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein.” Cell, vol. 98, no. 6, 1999, pp. 835–845., doi:10.1016/s0092-8674(00)81517-2. <ref/> | + | Specifically, there are several significant interaction patterns that stabilize adenosine recognition. RRM 1 and 2 makes significant interactions with the adenosine backbone, shown in Figure 3. Additionally, the adenosine stabilizes itself within the binding by intramolecular stacking interactions between adenosines. Through the extensive <scene name='78/781949/Lys_104_asp_105/1'>interactions with adenosine 2</scene>, the RRM specifies the position of adenosine 2, allowing it to make strong intramolecular stacking interactions with adenosine 1. As a result, adenosine 1 requires less contact with the RRM, as it is mostly stabilized by adenosine 2. Furthermore, some adenosines like adenosine 3 and adenosine 6 are stabilized by being sandwiched between aromatic and alipathic side chains. <scene name='78/781947/Interactions_with_a3/1'>Adenosine-3 sandwiching</scene> occurs between aromatic and alipathic side chains and is specified by Lysine 104, and <scene name='78/781947/Residues_interacting_with_a6/1'>Adenosine-6 sandwiching</scene> occurs similarly, but it is specified doubly by two residues, Trp-86 and Gln-88. <ref> Deo, Rahul C, et al. “Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein.” Cell, vol. 98, no. 6, 1999, pp. 835–845., doi:10.1016/s0092-8674(00)81517-2. <ref/>. |
===Translation Initiation=== | ===Translation Initiation=== | ||
| Line 39: | Line 39: | ||
== References == | == References == | ||
<references/> | <references/> | ||
| - | |||
| - | 1. Blatter, Markus, et al. “RNA Recognition Motifs: Boring? Not Quite.” Current Opinion in Structural Biology, vol. 18, no. 3, 2008, pp. 290–298., doi:10.1016/j.sbi.2008.04.002. | ||
| - | |||
| - | 2.De Melo Neto, Osvaldo P., et al. “Phosphorylation and Interactions Associated with the Control of the Leishmania Poly-A Binding Protein 1 (PABP1) Function during Translation Initiation.” RNA Biology, 23 Mar. 2018, pp. 1–17., doi:10.1080/15476286.2018.1445958. | ||
| - | |||
| - | 3.Deo, Rahul C, et al. “Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein.” Cell, vol. 98, no. 6, 1999, pp. 835–845., doi:10.1016/s0092-8674(00)81517-2. | ||
| - | |||
| - | 4.Gallie, Daniel R, and Renyi Liu. “Phylogenetic Analysis Reveals Dynamic Evolution of the Poly(A)-Binding Protein Gene Family in Plants.” BMC Evolutionary Biology, vol. 14, no. 1, 2014, doi:10.1186/s12862-014-0238-4. | ||
| - | |||
| - | 5.Gingras, Anne-Claude, et al. “eIF4 Initiation Factors: Effectors of MRNA Recruitment to Ribosomes and Regulators of Translation.” Annual Review of Biochemistry, vol. 68, no. 1, 1999, pp. 913–963., doi:10.1146/annurev.biochem.68.1.913. | ||
| - | |||
| - | 6.Imataka, H. “A Newly Identified N-Terminal Amino Acid Sequence of Human eIF4G Binds Poly(A)-Binding Protein and Functions in Poly(A)-Dependent Translation.” The EMBO Journal, vol. 17, no. 24, 1998, pp. 7480–7489., doi:10.1093/emboj/17.24.7480. | ||
| - | |||
| - | 7.Kahvejian, A. “Mammalian Poly(A)-Binding Protein Is a Eukaryotic Translation Initiation Factor, Which Acts via Multiple Mechanisms.” Genes & Development, vol. 19, no. 1, 2005, pp. 104–113., doi:10.1101/gad.1262905. | ||
| - | |||
| - | 8. Piron, M. “Rotavirus RNA-Binding Protein NSP3 Interacts with eIF4GI and Evicts the Poly(A) Binding Protein from eIF4F.” The EMBO Journal, vol. 17, no. 19, 1998, pp. 5811–5821., doi:10.1093/emboj/17.19.5811. | ||
Revision as of 17:32, 29 March 2018
Poly(A) binding protein - Homo sapiens
Structure
| |||||||||||