This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Poly(A) binding protein
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
=='''Poly(A) binding protein - ''Homo sapiens'''''== | =='''Poly(A) binding protein - ''Homo sapiens'''''== | ||
| - | |||
==Structure== | ==Structure== | ||
| Line 17: | Line 16: | ||
[[Image: Stacking_of_adenosines_with_beta_sheets.jpg |250 px|left|thumb|Figure 2: The specific weak intermolecular interactions between RNP1 and RNP2 and Adenosines. These interactions are the primary support of adenosine recognition by PABP and include mainly van der Waals interactions, hydrogen bonds, and stacking interactions. ]] | [[Image: Stacking_of_adenosines_with_beta_sheets.jpg |250 px|left|thumb|Figure 2: The specific weak intermolecular interactions between RNP1 and RNP2 and Adenosines. These interactions are the primary support of adenosine recognition by PABP and include mainly van der Waals interactions, hydrogen bonds, and stacking interactions. ]] | ||
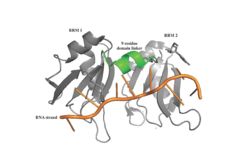
| - | A primary function of PABP is recognizing and interacting with the 3'poly (A) tail created in mRNA processing. As found by EMSA competition experiments, there is a minimum of 11-12 adenosines necessary in the poly (A) tail for the adenosine chain to bind to PABP with high affinity. However, for one biological assembly, a chain containing 9 adenosines sufficiently binds the assembly for crystallization and is shown in the biological assembly structure. The 4 RRM domains that are the primary interacting sites for the adenosine recognition exist as globular domains, each having four antiparallel β-strands and two α-helices. With the N-terminal to C-terminal motifs labeled as S1 to S4 for the β-strands and H1 to H2 for the α-helices, the strands are spatially arranged as S2-S3-S1-S4. Furthermore, there are two conserved sequences in each RRM, called RNP1 and 2. RNP 1 consists of a conserved sequence of 8 residues, while RNP2 consists of a conserved sequence of 6 residues. Much of the weak intermolecular interactions with adenosine from the RRMs occur from the <scene name='78/781946/Rnp1_rnp2_primary_a_binding/1'>RNP1 and RNP2</scene> conserved sequences, which correspond to the two central β-strands, with specific interactions shown in Figure 2.The support for adenosine recognition by the RRMs occurs as a type of binding trough with the sheets, primarily <scene name='78/781946/Rnp1_rnp2_primary_a_binding/1'>RNP1 and RNP2 forming the Primary Binding Trough</scene>, and the interstrand loop between β-strands 2 and 3 as well as the domain linker forming the <scene name='78/781946/Adenosine_binding_wall/1'> Adenosine Binding Wall</scene>. Additionally, the primary binding trough is stabilized by <scene name='78/781946/Rrm1_2_packing_intxn/2'>Stabilizing Packing Interactions of RRM1 RRM2 Binding Trough</scene>. | + | A primary function of PABP is recognizing and interacting with the [https://en.wikipedia.org/wiki/Polyadenylation 3'poly (A) tail] created in mRNA processing. As found by EMSA competition experiments, there is a minimum of 11-12 adenosines necessary in the poly (A) tail for the adenosine chain to bind to PABP with high affinity. However, for one biological assembly, a chain containing 9 adenosines sufficiently binds the assembly for crystallization and is shown in the biological assembly structure. The 4 RRM domains that are the primary interacting sites for the adenosine recognition exist as globular domains, each having four antiparallel β-strands and two α-helices. With the N-terminal to C-terminal motifs labeled as S1 to S4 for the β-strands and H1 to H2 for the α-helices, the strands are spatially arranged as S2-S3-S1-S4. Furthermore, there are two conserved sequences in each RRM, called RNP1 and 2. RNP 1 consists of a conserved sequence of 8 residues, while RNP2 consists of a conserved sequence of 6 residues. Much of the weak intermolecular interactions with adenosine from the RRMs occur from the <scene name='78/781946/Rnp1_rnp2_primary_a_binding/1'>RNP1 and RNP2</scene> conserved sequences, which correspond to the two central β-strands, with specific interactions shown in Figure 2.The support for adenosine recognition by the RRMs occurs as a type of binding trough with the sheets, primarily <scene name='78/781946/Rnp1_rnp2_primary_a_binding/1'>RNP1 and RNP2 forming the Primary Binding Trough</scene>, and the interstrand loop between β-strands 2 and 3 as well as the domain linker forming the <scene name='78/781946/Adenosine_binding_wall/1'> Adenosine Binding Wall</scene>. Additionally, the primary binding trough is stabilized by <scene name='78/781946/Rrm1_2_packing_intxn/2'>Stabilizing Packing Interactions of RRM1 RRM2 Binding Trough</scene>. |
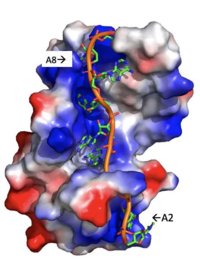
[[Image: Adenosine_backbone.png |200 px|right|thumb|Figure 3: Basic residues of RRM 1 and 2 (shown in blue) make stabilizing electrostatic interactions with the negatively charged adenosine phosphates. ]] | [[Image: Adenosine_backbone.png |200 px|right|thumb|Figure 3: Basic residues of RRM 1 and 2 (shown in blue) make stabilizing electrostatic interactions with the negatively charged adenosine phosphates. ]] | ||
Revision as of 17:54, 29 March 2018
Poly(A) binding protein - Homo sapiens
Structure
| |||||||||||