This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Ben Dawson/Sandbox1
From Proteopedia
| Line 5: | Line 5: | ||
== Background == | == Background == | ||
| - | The Human Poly-A Binding Protein (PABP) is a mRNA binding protein that binds to the 3’ Poly-A tail | + | The Human Poly-A Binding Protein (PABP) is a mRNA binding protein that binds to the 3’ Poly-A tail of mRNA prior to translation. Through extensive Adenosine recognition by the RNA Recognition Motifs (RRMs) of PABP, the protein is involved in three main functions: recognition of the 3’ Poly-A tail, mRNA stabilization, and, even further, eukaryotic translation initiation. The contributions of controlling gene expression via different families of PABPs is not yet fully understood. PABP families are divided into nuclear and cytoplasmic.5 PABP1, which is predominantly cytoplasmic, is often referred to as PABP because it is the only form of PABP that has been extensively studied in its role with mRNA translation and stability.5 |
| Line 11: | Line 11: | ||
The crystall structure was derived from X-ray Diffraction at 2.6Å (R-value: 23%). It is comprised of four RNA recognition motifs (RRMs), which are highly conserved RNA-binding domains.¹ The RRM in PABP is found in over two hundred families of proteins across species, indicating that it is ancient.¹ RRM1 and 2 are examined in this article. | The crystall structure was derived from X-ray Diffraction at 2.6Å (R-value: 23%). It is comprised of four RNA recognition motifs (RRMs), which are highly conserved RNA-binding domains.¹ The RRM in PABP is found in over two hundred families of proteins across species, indicating that it is ancient.¹ RRM1 and 2 are examined in this article. | ||
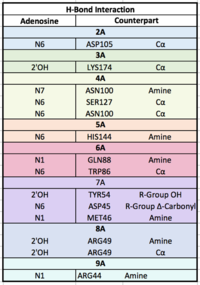
| - | Each RRM has four-stranded antiparallel beta sheet and two helices.¹ | + | Each RRM has a four-stranded antiparallel beta sheet and two alpha helices.¹ mRNA poly-adenosine recognition is due to the presence of the conserved residues within the beta-sheet surface², which forms a <scene name='78/782616/Trough2/1'>trough</scene>-like pocket for the mRNA to bind . The beta-sheet flooring present in PABP interacts with the 3’ mRNA tail via a combination of van der Waals, aromatic stacking, and Hydrogen bonding. Through these interactions, PABP binds to 3’ poly-adenosine tail with a KD of 2-7 nM.5 The structural elements highlighted consist of the RRM1/2 subunits, the linker domain, and the Poly-A mRNA binding trough. |
Revision as of 01:30, 2 April 2018
Human Poly A-Binding Protein (1CVJ)
| |||||||||||
References
1. Deo, Rahul C, et al. “Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein.” Cell 98:6. (1999) 835-845. Print.
2. Wang, Zuoren and Kiledjian, Megerditch. “The Poly(A)-Binding Protein and an mRNA Stability Protein Jointly Regulate an Endoribonuclease Activity.” Molecular and Cellular Biology 20.17 (2000): 6334–6341. Print.
3. “Oculopharyngeal Muscular Dystrophy.” NORD (National Organization for Rare Disorders), rarediseases.org/rare-diseases/oculopharyngeal-muscular-dystrophy/.
4. Richard, Pascale, et al. “Correlation between PABPN1 Genotype and Disease Severity in Oculopharyngeal Muscular Dystrophy.” Neurology, vol. 88, no. 4, 2016, pp. 359–365., doi:10.1212/wnl.0000000000003554.
5. Gorgoni, Barbra, and Gray, Nicola. “The Roles of Cytoplasmic Poly(A)-Binding Proteins in Regulating Gene Expression: A Developmental Perspective.” Briefings in Functional Genomics and Proteomics, vol. 3, no. 2, 1 Aug. 2004, pp. 125–141., doi:10.1093/bfgp/3.2.125.
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644