This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Alexis Neyman/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 9: | Line 9: | ||
== Structure == | == Structure == | ||
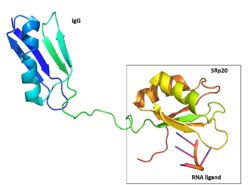
| - | [[Image:2D_RRM_and_RS_SRp20_with_fun_shapes3.png|250 px|right|thumb|Figure 1: SRp20 RRM and RS domains are shown]] The structure of SRp20 was determined by heteronuclear single quantum coherence ([https://en.wikipedia.org/wiki/Heteronuclear_single_quantum_coherence_spectroscopy HSQC]) NMR. The structure is composed of one RNA recognition motif (RRM) at the N-terminus and one Arg/Ser (AR) domain at the C-terminus where the Ser residues are phosphorylated<ref name="corbo">PMID:23685143</ref>. The RRM of SRp20 demonstrates the β1α1β2β3α2β3 topology seen in other [https://en.wikipedia.org/wiki/RNA_recognition_motif RRMs]. The role of the RRM region is to provide substrate specificity where SRp20 interacts with splicing enhancing sequences in mRNA. There have been no determined 3D structures of the RS domain thus it is unclear what its exact role is. However there have been some speculation that it might be involved in aiding protein-protein interactions in the spliceosome. It contains 164 amino acids, half belonging to the RRM and other half to the RS domain (Figure 1). SRp20 has a molecular weight of 19 kDA<ref name="corbo">PMID:23685143</ref>. | + | [[Image:2D_RRM_and_RS_SRp20_with_fun_shapes3.png|250 px|right|thumb|Figure 1: SRp20 RRM and RS domains are shown]] The structure of SRp20 was determined by heteronuclear single quantum coherence ([https://en.wikipedia.org/wiki/Heteronuclear_single_quantum_coherence_spectroscopy HSQC]) NMR. The structure is composed of one RNA recognition motif (RRM) at the N-terminus and one Arg/Ser (AR) domain at the C-terminus where the Ser residues are phosphorylated<ref name="corbo">PMID:23685143</ref>. The RRM of SRp20 demonstrates the β1α1β2β3α2β3 topology seen in other [https://en.wikipedia.org/wiki/RNA_recognition_motif RRMs]. The role of the RRM region is to provide substrate specificity where SRp20 interacts with splicing enhancing sequences in mRNA. There have been no determined 3D structures of the RS domain thus it is unclear what its exact role is. However, there have been some speculation that it might be involved in aiding protein-protein interactions in the spliceosome. It contains 164 amino acids, half belonging to the RRM and other half to the RS domain (Figure 1). SRp20 has a molecular weight of 19 kDA<ref name="corbo">PMID:23685143</ref>. |
=== Poor Solubility Problem === | === Poor Solubility Problem === | ||
| Line 17: | Line 17: | ||
1H-15N HSQC results showed a large hydrophobic β-sheet on the RRM binding to the RNA with all four bases interacting with one of the four aromatic residues via hydrophobic interactions <ref name="Hargous">PMID:17036044</ref>. [https://en.wikipedia.org/wiki/Beta_hairpin β-hairpin] amino acids are hydrogen bonded to bases on nucleic acid targets <ref name="Clery">PMID:18515081</ref>. This suggests that the β-hairpin plays a role in SRp20 selectivity for specific ligands. The researchers used a smaller peptide chain to reduce the NMR broadening seen with longer peptides (allowing for structure determination), with the consequence of reduced binding affinity. | 1H-15N HSQC results showed a large hydrophobic β-sheet on the RRM binding to the RNA with all four bases interacting with one of the four aromatic residues via hydrophobic interactions <ref name="Hargous">PMID:17036044</ref>. [https://en.wikipedia.org/wiki/Beta_hairpin β-hairpin] amino acids are hydrogen bonded to bases on nucleic acid targets <ref name="Clery">PMID:18515081</ref>. This suggests that the β-hairpin plays a role in SRp20 selectivity for specific ligands. The researchers used a smaller peptide chain to reduce the NMR broadening seen with longer peptides (allowing for structure determination), with the consequence of reduced binding affinity. | ||
| - | The ligand used was <scene name='78/781963/Looking_at_the_ligand/1'>CAUC</scene>. The conformation of U3 and C4 shows that U3 bulges out while C4 partially stacks over A2. Interactions with the RRM that | + | The ligand used was <scene name='78/781963/Looking_at_the_ligand/1'>CAUC</scene>. The conformation of U3 and C4 shows that U3 bulges out while C4 partially stacks over A2. Interactions with the RRM that were noticed were that <scene name='78/781963/C1_and_tyr_13/3'>C1 stacks with Tyr 13</scene> in β1 and <scene name='78/781963/A2_phe_50/2'>A2 stacks with Phe 50</scene> in β3. These aromatic side chains form hydrophobic interactions with the ligand when stacked (Figure 3). Also, the residue <scene name='78/781963/C1_a2_phe48/2'>F48 inserts between the sugar rings of C1 and A2</scene>. <scene name='78/781963/C1_binding_pocket3/1'>C1 is recognized definitively by the RRM</scene>. The amino proton of C1 hydrogen bonds with the carbonyl oxygen of Leu 80 and the side-chain carbonyl oxygen of Glu 79. The N3 of C1 hydrogen bonds with the amide of Asn 82, and the O2 of C1 hydrogen bonds with the hydroxyl group of Ser 81<ref name="Hargous">PMID:17036044</ref>. |
It was also noted that <scene name='78/781963/A2_syn_conformation/1'>A2</scene> adopts an unusual syn conformation. U3 interacts with <scene name='78/781963/U3_hydrophobic_interactions/2'>Phe 48, Trp 40, Ala 42,</scene> and with the β2-3 loop of the RRM. These residues are all hydrophobic, offering a large hydrophobic surface that helps bind the ligand, as well as prevents the solvent from binding. Additionally, C4 is maintained in its position by a <scene name='78/781963/C4_a2_h_bond/1'>hydrogen bond between C4 amino group and the A2 2’ oxygen</scene> <ref name="Hargous">PMID:17036044</ref>. [[Image:Figure_4_C1_and_A2_interactions_Edited2.png|300 px|left|thumb|Figure 3: C1 and A2 on the RNA ligand interacting with hydrophobic residues (Tyr 13, Phe 50, Phe 48) in the RRM domain of the SRp20 protein. Image created using ''Pymol'']] | It was also noted that <scene name='78/781963/A2_syn_conformation/1'>A2</scene> adopts an unusual syn conformation. U3 interacts with <scene name='78/781963/U3_hydrophobic_interactions/2'>Phe 48, Trp 40, Ala 42,</scene> and with the β2-3 loop of the RRM. These residues are all hydrophobic, offering a large hydrophobic surface that helps bind the ligand, as well as prevents the solvent from binding. Additionally, C4 is maintained in its position by a <scene name='78/781963/C4_a2_h_bond/1'>hydrogen bond between C4 amino group and the A2 2’ oxygen</scene> <ref name="Hargous">PMID:17036044</ref>. [[Image:Figure_4_C1_and_A2_interactions_Edited2.png|300 px|left|thumb|Figure 3: C1 and A2 on the RNA ligand interacting with hydrophobic residues (Tyr 13, Phe 50, Phe 48) in the RRM domain of the SRp20 protein. Image created using ''Pymol'']] | ||
| Line 34: | Line 34: | ||
== Relationship to 9G8 == | == Relationship to 9G8 == | ||
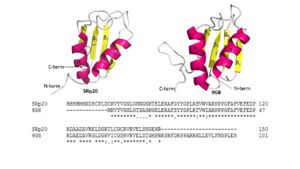
| - | [[Image:Combined_SRp20_and_9G8_Image.jpg|300 px|left|thumb|Figure 4: Comparing SRp20 and 9G8 RRMs and sequence alignments. Structural images created using ''Pymol'']] <scene name='78/781963/Rrm_motif/1'>SRp20</scene> and splicing factor <scene name='78/781963/9g8_rrm/1'>9G8</scene> are both sequence specific RNA binding proteins (Figure 4) and are the smallest members of the Serine-and-Arginine Rich (SR) protein family. Both RNA Recognition Motifs (RRMs) have a similar βαββαβ topology. SRp20 and 9G8 are 80% identical. The sequence alignment shows the alignment of the RRMs of SRp20 and 9G8 <ref name="Hargous">PMID:17036044</ref>. SRp20 binds pyrimidine rich areas while 9G8 binds purine rich areas.This difference in binding comes from the fact that 9G8 has a [https://en.wikipedia.org/wiki/Zinc_finger zinc knuckle] that recognizes GAC triplets <ref name="Cava">PMID:10094314 </ref>. 9G8s RRM is followed by a zinc knuckle and then the SR domain whereas SRp20s RRM is followed directly by the SR domain. When 9G8 lacks a zinc knuckle, it binds pyrimidine-rich sequences like SRp20 <ref name="Hargous">PMID:17036044</ref>. The zinc knuckle of 9G8 contains glycine residues at positions 5 and 8 and charged residues at positions 6 and 13 that are highly conserved <ref name="Cava">PMID:10094314 </ref>. Due to the poor solubility problem, a structure for the zinc knuckle of 9G8 is not available to show in an image. | + | [[Image:Combined_SRp20_and_9G8_Image.jpg|300 px|left|thumb|Figure 4: Comparing SRp20 and 9G8 RRMs and sequence alignments. Structural images created using ''Pymol'']] <scene name='78/781963/Rrm_motif/1'>SRp20</scene> and splicing factor <scene name='78/781963/9g8_rrm/1'>9G8</scene> are both sequence specific RNA binding proteins (Figure 4) and are the smallest members of the Serine-and-Arginine Rich (SR) protein family. Both RNA Recognition Motifs (RRMs) have a similar βαββαβ topology. SRp20 and 9G8 are 80% identical. The sequence alignment shows the alignment of the RRMs of SRp20 and 9G8 <ref name="Hargous">PMID:17036044</ref>(Figure 4). SRp20 binds pyrimidine rich areas while 9G8 binds purine rich areas.This difference in binding comes from the fact that 9G8 has a [https://en.wikipedia.org/wiki/Zinc_finger zinc knuckle] that recognizes GAC triplets <ref name="Cava">PMID:10094314 </ref>. 9G8s RRM is followed by a zinc knuckle and then the SR domain whereas SRp20s RRM is followed directly by the SR domain. When 9G8 lacks a zinc knuckle, it binds pyrimidine-rich sequences like SRp20 <ref name="Hargous">PMID:17036044</ref>. The zinc knuckle of 9G8 contains glycine residues at positions 5 and 8 and charged residues at positions 6 and 13 that are highly conserved <ref name="Cava">PMID:10094314 </ref>. Due to the poor solubility problem, a structure for the zinc knuckle of 9G8 is not available to show in an image. |
== Disease == | == Disease == | ||
Revision as of 16:12, 17 April 2018
Biological Structure of SRp20
| |||||||||||