This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Ethan Kitt/Sandbox 1
From Proteopedia
| Line 6: | Line 6: | ||
==Structure== | ==Structure== | ||
| - | <StructureSection load='1B7F' size='350' frame='true' side='right' caption='PABP' scene='78/782614/Structure_scene_color_scheme/1'> The crystal structure PABP was derived from X-ray Diffraction at 2.6Å (R-value: 23%). The subunits of PABP, RRM1 and RRM2, are examined in this article as the ''in vivo'' form seen in biological assembly 1 (via PDB). The protein has a homopolymeric structure, containing four RNA recognition motifs (RRMs), which are conserved. <ref name="Structure and Function">Kühn, Uwe and Elmar, Wahle. “Structure and Function of Poly(a) Binding Proteins.” Bba - Gene Structure & Expression, vol. 1678, no. 2/3, 2004. </ref> <scene name='78/782616/Rrm1_only/2'>RRM1</scene> and <scene name='78/782616/Rrm2_only/2'>RRM2</scene> are N-terminal domains that are connected by a <scene name='78/782616/Linker/3'>linker</scene>.<ref name="Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein">Deo, Rahul C, et al. “Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein.” Cell 98:6. (1999) 835-845. Print. </ref> Opposed to their counterparts, RRM3 and RRM4 bind Poly (A) RNA less tightly than RRM1 and RRM2.<ref name="Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein">Deo, Rahul C, et al. “Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein.” Cell 98:6. (1999) 835-845. Print. </ref> | + | <StructureSection load='1B7F' size='350' frame='true' side='right' caption='PABP' scene='78/782614/Structure_scene_color_scheme/1'> |
| + | The crystal structure PABP was derived from X-ray Diffraction at 2.6Å (R-value: 23%). The subunits of PABP, RRM1 and RRM2, are examined in this article as the ''in vivo'' form seen in biological assembly 1 (via PDB). The protein has a homopolymeric structure, containing four RNA recognition motifs (RRMs), which are conserved. <ref name="Structure and Function">Kühn, Uwe and Elmar, Wahle. “Structure and Function of Poly(a) Binding Proteins.” Bba - Gene Structure & Expression, vol. 1678, no. 2/3, 2004. </ref> <scene name='78/782616/Rrm1_only/2'>RRM1</scene> and <scene name='78/782616/Rrm2_only/2'>RRM2</scene> are N-terminal domains that are connected by a <scene name='78/782616/Linker/3'>linker</scene>.<ref name="Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein">Deo, Rahul C, et al. “Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein.” Cell 98:6. (1999) 835-845. Print. </ref> Opposed to their counterparts, RRM3 and RRM4 bind Poly (A) RNA less tightly than RRM1 and RRM2.<ref name="Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein">Deo, Rahul C, et al. “Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein.” Cell 98:6. (1999) 835-845. Print. </ref> | ||
===RRMs=== | ===RRMs=== | ||
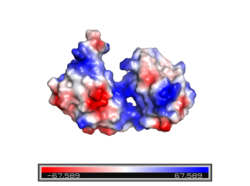
[[Image:ElectroPOSITIVE.png|250 px|left|thumb|Figure 2. Electropositive binding cleft. Electropositive region shown in blue and electronegative region shown in red.]] The sex-lethal protein (Sxl) is 354 amino acid residues long. It is composed of two highly conserved regions called <scene name='78/782600/Rbd1_and_rbd2/4'>RNA recognition motifs</scene> (RRMs) that function as a monomeric unit. Each RRM is approximately 90 amino acids long with a four stranded sheet <scene name='78/782600/Opening_image/3'>β-pleated</scene> and two <scene name='78/782600/Alpha_helices/1'>α-helices</scene>. The β-pleated sheets from each RRM interact with the RNA ligand, while the α-helices interact with each other to shape the protein <ref name="Clery"/>. The RRMs interact at <scene name='78/782600/Inter-domain_interactions/5'>two places</scene>: between the side chain of Lys 197 and the main chain carbonyl of Val 238 and between the side chains of Tyr 131 and Gln 239<ref name="Handa" />. The two RRMs are connected via an interdomain linker. The linker often forms a short 3<sub>10</sub> helix from Gly205 to Thr211.The interaction between the RRMs and the ligand is facilitated by the v-shaped <scene name='78/782600/Binding_cleft/7'>binding cleft</scene> formed by the β-pleated sheet of each RRM. The v-shaped cleft is strongly electropositive which also assists in ligand binding <ref name="Handa" /> (Figure 2). The presence of two RRMs increases RNA binding specificity by allowing for an elongated and continuous binding site <ref name="Clery">DOI 10.1016/j.sbi.2008.04.002</ref>. | [[Image:ElectroPOSITIVE.png|250 px|left|thumb|Figure 2. Electropositive binding cleft. Electropositive region shown in blue and electronegative region shown in red.]] The sex-lethal protein (Sxl) is 354 amino acid residues long. It is composed of two highly conserved regions called <scene name='78/782600/Rbd1_and_rbd2/4'>RNA recognition motifs</scene> (RRMs) that function as a monomeric unit. Each RRM is approximately 90 amino acids long with a four stranded sheet <scene name='78/782600/Opening_image/3'>β-pleated</scene> and two <scene name='78/782600/Alpha_helices/1'>α-helices</scene>. The β-pleated sheets from each RRM interact with the RNA ligand, while the α-helices interact with each other to shape the protein <ref name="Clery"/>. The RRMs interact at <scene name='78/782600/Inter-domain_interactions/5'>two places</scene>: between the side chain of Lys 197 and the main chain carbonyl of Val 238 and between the side chains of Tyr 131 and Gln 239<ref name="Handa" />. The two RRMs are connected via an interdomain linker. The linker often forms a short 3<sub>10</sub> helix from Gly205 to Thr211.The interaction between the RRMs and the ligand is facilitated by the v-shaped <scene name='78/782600/Binding_cleft/7'>binding cleft</scene> formed by the β-pleated sheet of each RRM. The v-shaped cleft is strongly electropositive which also assists in ligand binding <ref name="Handa" /> (Figure 2). The presence of two RRMs increases RNA binding specificity by allowing for an elongated and continuous binding site <ref name="Clery">DOI 10.1016/j.sbi.2008.04.002</ref>. | ||
Revision as of 18:09, 20 April 2018
Contents |
Human Poly(A) Binding Protein (1CVJ)
Background
The Human Poly(A) Binding Protein (PABP) was discovered in 1973 by the use of a sedimentation profile detailing the RNase digestion differentiated the PABP protein. [1] Attempts to purify the 75 kDa protein then followed. In 1983, then considered “poly(A)-organizing protein,” was determined and purified by molecular weight, ligand-binding affinity, and amounts found in cytoplasmic portions of cell with ability to bind to free poly(A). [2]
Structure
| |||||||||||
Autoregulation
Sxl controls its own levels of expression via positive and negative autoregulation. Sxl binds its own pre-mRNA transcript in a similar manner as its downstream targets, Tra and Msl2. Through binding to its recognition element, Sxl causes a 3’ splice site to be skipped. Alternative splicing occurs utilizing a 3’ splice site further downstream, cleaving out a premature stop codon within Exon 3 and preventing truncation and inactivation[7]. This is a pathway of positive autoregulation, as functional Sxl protein must be present to cause proper processing of the pre-mRNA. The negative autoregulation pathway of Sxl proceeds via repression of its own translation. The Sxl transcript contains the target polyuridine sequence within its 3’UTR. Sxl binds this target, and blocks translation[7]. Negative autoregulation allows maintenance of a stable and standard Sxl protein concentration. An excess of Sxl increases the degree of translation repression because more Sxl protein are present to potentially bind at the 3’UTR, while a shortage allows for more unrepressed translation.
Mutation
In fruit flies, dosage compensation is achieved by hyperexpression of the single X chromosome found in the male genome. The single X chromosome is hyperexpressed and regulated by a group of genes known as the Male-specific lethal genes. The genes involve work together to produce a complex known as the Male-specific lethal complex (Msl complex). One gene that is important in forming this complex is the gene that codes for the Msl-2 protein. Without this protein, the Msl complex cannot form [8]. Males exclusively make the Msl-2 protein and therefore exclusively create the Msl complex. Female cannot make the complex because the Sxl protein prevents the production of the Msl-2 protein. It prevents the protein formation by alternatively splicing of an intron near the 5’ UTR. In males this intron is included and in females it is not. [8] Mutations of the Sxl protein can affect the formation of this complex and lead to death for females—hence the name sex lethal. If the Sxl gene is mutated in females, the formation of the Msl-2 complex will not be inhibited. If there is a functional Msl-2 complex in females then both of the X chromosomes will be hyperexpressed [8] The overexpression of genes on the X chromosome is a fatal phenomenon ultimately caused by a mutation of the Sxl protein. Although a mutation to the Sxl gene is harmful to females, it has no effect on males because the functional Sxl gene is not expressed—a mutation in males has no phenotypic effect.
References
- ↑ Blobel, Gunter. “A Protein of Molecular Weight 78,000 Bound to the Polyadenylate Region of Eukaryotic Messenger Rnas.” Proceedings of the National Academy of Sciences of the United States of America, vol. 70, no. 3, 1973, pp. 924–8.
- ↑ Baer, Bradford W. and Kornberg, Roger D. "The Protein Responsible for the Repeating Structure of Cytoplasmic Poly(A)-Ribonucleoprotein." The Journal of Cell Biology, vol. 96, no. 3, Mar. 1983, pp. 717-721. EBSCOhost.
- ↑ Kühn, Uwe and Elmar, Wahle. “Structure and Function of Poly(a) Binding Proteins.” Bba - Gene Structure & Expression, vol. 1678, no. 2/3, 2004.
- ↑ 4.0 4.1 Deo, Rahul C, et al. “Recognition of Polyadenylate RNA by the Poly(A)-Binding Protein.” Cell 98:6. (1999) 835-845. Print.
- ↑ 5.0 5.1 Clery A, Blatter M, Allain FH. RNA recognition motifs: boring? Not quite. Curr Opin Struct Biol. 2008 Jun;18(3):290-8. doi: 10.1016/j.sbi.2008.04.002. PMID:18515081 doi:http://dx.doi.org/10.1016/j.sbi.2008.04.002
- ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedHanda - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedBlack - ↑ 8.0 8.1 8.2 Penalva L, Sanchez L. RNA Binding Protein Sex-Lethal (Sxl) and Control of Drosophila Sex Determination and Dosage Compensation. Microbiol Mol Biol Rev.;67(3):343-356. doi: 10.1128/MMBR.67.3.343–359.2003