This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Nuclear polyadenylated RNA-binding protein
From Proteopedia
(Difference between revisions)
| Line 8: | Line 8: | ||
=Structure= | =Structure= | ||
==General Features== | ==General Features== | ||
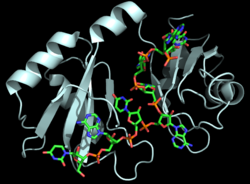
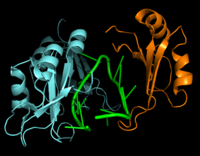
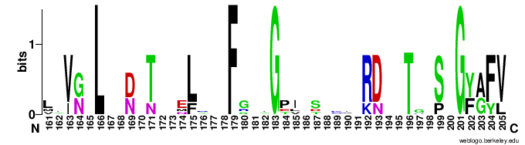
| - | Hrp1 is a single-stranded [https://en.wikipedia.org/wiki/RNA-binding_protein RNA-binding protein] composed of two RNP-type [https://en.wikipedia.org/wiki/RNA_recognition_motif RNA-binding domains (RBDs)] arranged in tandem with a typical ßαßßαß architecture <ref name="GM3H"/>. The two RBDs have similar topolgies, both containing a central [https://en.wikipedia.org/wiki/Beta_sheet antiparallel] four-stranded <scene name='78/783765/Beta_sheet/1'>ß-sheet</scene> with two [https://en.wikipedia.org/wiki/Alpha_helix α-helices] running across one face <ref name="GM3H"/>. The two RBDs associate to form a deep and positively charged <scene name='78/781960/Hrp1-rna_interface_surface/2'>cleft</scene>, which constitutes the binding site for the RNA molecule <ref name="GM3H"/>. | + | Hrp1 is a single-stranded [https://en.wikipedia.org/wiki/RNA-binding_protein RNA-binding protein] composed of two RNP-type [https://en.wikipedia.org/wiki/RNA_recognition_motif RNA-binding domains (RBDs)] arranged in tandem with a typical ßαßßαß architecture <ref name="GM3H"/>. The two RBDs have similar topolgies, both containing a central [https://en.wikipedia.org/wiki/Beta_sheet antiparallel] four-stranded <scene name='78/783765/Beta_sheet/1'>ß-sheet</scene> with two [https://en.wikipedia.org/wiki/Alpha_helix α-helices] running across one face <ref name="GM3H"/>. The β-strands of each βαβ domain are linked via hydrogen bonding between conserved residues, <scene name='78/783765/L166_g201/2'>Leu166 and Gly201</scene>. The two RBDs associate to form a deep and positively charged <scene name='78/781960/Hrp1-rna_interface_surface/2'>cleft</scene>, which constitutes the binding site for the RNA molecule <ref name="GM3H"/>. |
==Hrp1-RNA Interactions== | ==Hrp1-RNA Interactions== | ||
Revision as of 14:49, 24 April 2018
| |||||||||||
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 1.17 1.18 1.19 1.20 1.21 Perez-Canadillas JM. Grabbing the message: structural basis of mRNA 3'UTR recognition by Hrp1. EMBO J. 2006 Jul 12;25(13):3167-78. Epub 2006 Jun 22. PMID:16794580
- ↑ 2.0 2.1 2.2 2.3 2.4 Leeper TC, Qu X, Lu C, Moore C, Varani G. Novel protein-protein contacts facilitate mRNA 3'-processing signal recognition by Rna15 and Hrp1. J Mol Biol. 2010 Aug 20;401(3):334-49. Epub 2010 Jun 19. PMID:20600122 doi:10.1016/j.jmb.2010.06.032
- ↑ Kessler MM, Henry MF, Shen E, Zhao J, Gross S, Silver PA, Moore CL. Hrp1, a sequence-specific RNA-binding protein that shuttles between the nucleus and the cytoplasm, is required for mRNA 3'-end formation in yeast. Genes Dev. 1997 Oct 1;11(19):2545-56. PMID:9334319
- ↑ Clery A, Blatter M, Allain FH. RNA recognition motifs: boring? Not quite. Curr Opin Struct Biol. 2008 Jun;18(3):290-8. doi: 10.1016/j.sbi.2008.04.002. PMID:18515081 doi:http://dx.doi.org/10.1016/j.sbi.2008.04.002
Proteopedia Page Contributors and Editors (what is this?)
Cory A. Wuerch, Matthew Douglas Moore, Savannah Davis, Michal Harel, Jaime Prilusky