This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Christian Fjeld/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
== General Description == | == General Description == | ||
<StructureSection load='4aq7' frame='true' size='340' side='right' caption='LARS (E coli) ternary complex with tRNAleu and leucyl adenylate analogue' scene='78/786656/Lars/2'> | <StructureSection load='4aq7' frame='true' size='340' side='right' caption='LARS (E coli) ternary complex with tRNAleu and leucyl adenylate analogue' scene='78/786656/Lars/2'> | ||
| - | <scene name='78/786656/Total/1'>Leucyl tRNA synthetase (LARS)</scene> is a 97 kDa, class IA aminoacyl-tRNA synthetase (ARS) that catalyzes the ligation of leucine with tRNA<sup>leu</sup> in an ATP dependent mechanism. LARS is a cytoplasmic enzyme that is found as part of the multisynthetase complex in eukaryotes<ref>doi: 10.1007/978-3-319-46503-6_18</ref>. The multi-synthetase complex contains glutamylprolyl-tRNA synthetase (EPRS), isoleucyl (IARS), leucyl (LARS), glutaminyl (GARS), methionyl (MARS), lysyl (KARS), arginyl (RARS), and aspartyl (DARS) tRNA synthetases as well as p18, p38 and p43<ref>doi: 10.1016/S0006-291X(03)00485-6</ref>. | + | <scene name='78/786656/Total/1'>Leucyl tRNA synthetase (LARS)</scene> is a 97 kDa, class IA aminoacyl-tRNA synthetase (ARS) that catalyzes the ligation of leucine with tRNA<sup>leu</sup> in an ATP dependent mechanism. ARS are essential to all life as they charge amino acids onto cognate tRNAs in preparation for protein translation. This is step is a potential source of error in interpreting the genetic code as mischarged tRNAs will not be recognized during protein synthesis and could lead to nonfunctional proteins. As such, ARS have a high specificity for both tRNA and amino acids and contain an editing domain capable of hydrolyzing mischarged tRNA. |
| + | |||
| + | LARS is a cytoplasmic enzyme that is found as part of the multisynthetase complex in eukaryotes<ref>doi: 10.1007/978-3-319-46503-6_18</ref>. The multi-synthetase complex contains glutamylprolyl-tRNA synthetase (EPRS), isoleucyl (IARS), leucyl (LARS), glutaminyl (GARS), methionyl (MARS), lysyl (KARS), arginyl (RARS), and aspartyl (DARS) tRNA synthetases as well as p18, p38 and p43<ref>doi: 10.1016/S0006-291X(03)00485-6</ref>. | ||
Multisynthetase complexes have also been seen in some archaea such as ''Thermococcus kodakarensis'' although the composition of the complex is not the same as eukaryotes<ref>doi: 10.1016/j.febslet.2012.05.039</ref>. | Multisynthetase complexes have also been seen in some archaea such as ''Thermococcus kodakarensis'' although the composition of the complex is not the same as eukaryotes<ref>doi: 10.1016/j.febslet.2012.05.039</ref>. | ||
LARS has been shown to be involved with the mTOR pathways as a sensor of leucine levels within the cell<ref>doi: 10.1016/j.cell.2012.02.044</ref>. | LARS has been shown to be involved with the mTOR pathways as a sensor of leucine levels within the cell<ref>doi: 10.1016/j.cell.2012.02.044</ref>. | ||
| + | |||
| + | === Disease === | ||
Mutations in LARS2, the mitochondrial version of the enzyme, have been linked to Perrault syndrome characterized by premature ovarian failure in females and progressive hearing loss in both sexes<ref>doi: 10.1016/j.ajhg.2013.03.007</ref> | Mutations in LARS2, the mitochondrial version of the enzyme, have been linked to Perrault syndrome characterized by premature ovarian failure in females and progressive hearing loss in both sexes<ref>doi: 10.1016/j.ajhg.2013.03.007</ref> | ||
| Line 10: | Line 14: | ||
=== Catalytic Domain === | === Catalytic Domain === | ||
| - | The <scene name='78/786656/Catalytic/1'>catalytic domain</scene> is responsible for the two step process of charging leucine on to tRNA<sup>leu</sup>. First, ATP and leucine are bound and AMP is transfered to the backbone carboxylic acid of leucine with the release of a pyrophosphate. Second, tRNA<sup>leu</sup> is bound with the leucyl adenylate and leucine is transfered to either the 2 | + | |
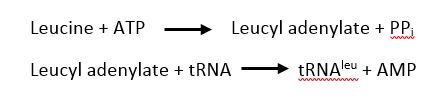
| + | The <scene name='78/786656/Catalytic/1'>catalytic domain</scene> is responsible for the two step process of charging leucine on to tRNA<sup>leu</sup>. First, ATP and leucine are bound and AMP is transfered to the backbone carboxylic acid of leucine with the release of a pyrophosphate. Second, tRNA<sup>leu</sup> is bound with the leucyl adenylate and leucine is transfered to either the 2' OH of the 3' terminal adenine with the release of AMP<ref>doi: 10.1038/nsmb.2317</ref>. | ||
[[Image:Scheme.JPG]] | [[Image:Scheme.JPG]] | ||
| + | |||
| + | LARS, like all class I synthetases, is characterized by a Rossmann-fold catalytic domain with a central parallel β-sheet with α-helices on both faces. The active site catalyzes both the formation of the leucyl adenylate intermediate and the subsequent charging of leucine onto the terminal acceptor arm of the tRNA. | ||
<ref>doi: 10.1093/emboj/19.10.2351</ref> | <ref>doi: 10.1093/emboj/19.10.2351</ref> | ||
| - | <ref>doi: 10.1038/nsmb.2317</ref> | ||
=== Editing Domain === | === Editing Domain === | ||
| + | |||
The <scene name='78/786656/Editing/1'>editing domain</scene> of LARS is responsible for editing mischarged tRNA and ensuring translational fidelity. | The <scene name='78/786656/Editing/1'>editing domain</scene> of LARS is responsible for editing mischarged tRNA and ensuring translational fidelity. | ||
<ref>doi: 10.1016/j.jmb.2009.04.073</ref> | <ref>doi: 10.1016/j.jmb.2009.04.073</ref> | ||
| Line 23: | Line 30: | ||
=== Anticodon Binding Domain === | === Anticodon Binding Domain === | ||
| + | |||
The <scene name='78/786656/Anticodon/1'>anticodon binding domain</scene> is essential for the fidelity of ARSs. However, there are numerous anticodons that correspond to leucine including AAU, GUA, and GUG | The <scene name='78/786656/Anticodon/1'>anticodon binding domain</scene> is essential for the fidelity of ARSs. However, there are numerous anticodons that correspond to leucine including AAU, GUA, and GUG | ||
=== Zinc Binding Domain === | === Zinc Binding Domain === | ||
| + | |||
The <scene name='78/786656/Zn1/1'>zinc binding domain</scene> | The <scene name='78/786656/Zn1/1'>zinc binding domain</scene> | ||
=== Leucine Specificity Domain === | === Leucine Specificity Domain === | ||
| + | |||
The <scene name='78/786656/Ls/1'>leucine specificity domain</scene> | The <scene name='78/786656/Ls/1'>leucine specificity domain</scene> | ||
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
| + | |||
A <scene name='78/786656/Conservation/2'>map of the conserved residues</scene> | A <scene name='78/786656/Conservation/2'>map of the conserved residues</scene> | ||
{{Template:ColorKey_ConSurf}} | {{Template:ColorKey_ConSurf}} | ||
| - | == Disease == | ||
</StructureSection> | </StructureSection> | ||
| + | |||
== 3D Structure of LARS == | == 3D Structure of LARS == | ||
| + | |||
Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | ||
{{#tree:id=OrganizedByTopic|openlevels=0| | {{#tree:id=OrganizedByTopic|openlevels=0| | ||
Revision as of 02:17, 2 May 2018
General Description
| |||||||||||
3D Structure of LARS
Updated on 02-May-2018
References
- ↑ Mirande M. The Aminoacyl-tRNA Synthetase Complex. Subcell Biochem. 2017;83:505-522. doi: 10.1007/978-3-319-46503-6_18. PMID:28271488 doi:http://dx.doi.org/10.1007/978-3-319-46503-6_18
- ↑ Han JM, Kim JY, Kim S. Molecular network and functional implications of macromolecular tRNA synthetase complex. Biochem Biophys Res Commun. 2003 Apr 18;303(4):985-93. doi: , 10.1016/s0006-291x(03)00485-6. PMID:12684031 doi:http://dx.doi.org/10.1016/s0006-291x(03)00485-6
- ↑ Raina M, Elgamal S, Santangelo TJ, Ibba M. Association of a multi-synthetase complex with translating ribosomes in the archaeon Thermococcus kodakarensis. FEBS Lett. 2012 Jul 30;586(16):2232-8. doi: 10.1016/j.febslet.2012.05.039. Epub, 2012 Jun 7. PMID:22683511 doi:http://dx.doi.org/10.1016/j.febslet.2012.05.039
- ↑ Han JM, Jeong SJ, Park MC, Kim G, Kwon NH, Kim HK, Ha SH, Ryu SH, Kim S. Leucyl-tRNA synthetase is an intracellular leucine sensor for the mTORC1-signaling pathway. Cell. 2012 Apr 13;149(2):410-24. doi: 10.1016/j.cell.2012.02.044. Epub 2012 Mar, 15. PMID:22424946 doi:http://dx.doi.org/10.1016/j.cell.2012.02.044
- ↑ Pierce SB, Gersak K, Michaelson-Cohen R, Walsh T, Lee MK, Malach D, Klevit RE, King MC, Levy-Lahad E. Mutations in LARS2, encoding mitochondrial leucyl-tRNA synthetase, lead to premature ovarian failure and hearing loss in Perrault syndrome. Am J Hum Genet. 2013 Apr 4;92(4):614-20. doi: 10.1016/j.ajhg.2013.03.007. Epub, 2013 Mar 28. PMID:23541342 doi:http://dx.doi.org/10.1016/j.ajhg.2013.03.007
- ↑ Palencia A, Crepin T, Vu MT, Lincecum TL Jr, Martinis SA, Cusack S. Structural dynamics of the aminoacylation and proofreading functional cycle of bacterial leucyl-tRNA synthetase. Nat Struct Mol Biol. 2012 Jun 10. doi: 10.1038/nsmb.2317. PMID:22683997 doi:10.1038/nsmb.2317
- ↑ Cusack S, Yaremchuk A, Tukalo M. The 2 A crystal structure of leucyl-tRNA synthetase and its complex with a leucyl-adenylate analogue. EMBO J. 2000 May 15;19(10):2351-61. PMID:10811626 doi:10.1093/emboj/19.10.2351
- ↑ Seiradake E, Mao W, Hernandez V, Baker SJ, Plattner JJ, Alley MR, Cusack S. Crystal structures of the human and fungal cytosolic Leucyl-tRNA synthetase editing domains: A structural basis for the rational design of antifungal benzoxaboroles. J Mol Biol. 2009 Jul 10;390(2):196-207. Epub 2009 May 6. PMID:19426743 doi:10.1016/j.jmb.2009.04.073
- ↑ Liu Y, Liao J, Zhu B, Wang ED, Ding J. Crystal structures of the editing domain of Escherichia coli leucyl-tRNA synthetase and its complexes with Met and Ile reveal a lock-and-key mechanism for amino acid discrimination. Biochem J. 2006 Mar 1;394(Pt 2):399-407. PMID:16277600 doi:10.1042/BJ20051249